Introduction
Oncologists face a persistent monitoring gap: standard imaging-based response criteria like RECIST can only detect tumor size changes weeks or months after treatment begins. RECIST assessments typically occur every 6–8 weeks, yet tumors can develop resistance, shed resistant clones, or regrow at the molecular level long before anatomical changes become visible.
Imaging also cannot capture minimal residual disease (MRD) after curative-intent surgery. MRD refers to microscopic tumor remnants that drive relapse but remain undetectable on CT and MRI.
Circulating tumor DNA (ctDNA) analysis via liquid biopsy addresses this gap directly — offering a real-time, blood-based window into tumor dynamics. Studies show that ctDNA-based molecular progression can precede radiographic progression by a median of 4.8 months.
This article covers what ctDNA analysis is, why it has become central to treatment monitoring in precision oncology, how the lab workflow operates in practice, and which pre-analytical factors determine whether results are clinically reliable.
TL;DR
- ctDNA is tumor-derived DNA circulating in blood, carrying tumor-specific mutations and distinctive fragmentation patterns
- Detects molecular changes weeks to months before imaging, enabling earlier treatment decisions
- Key applications: treatment response confirmation, MRD detection, resistance mutation surveillance
- Extraction quality is the make-or-break pre-analytical step: poor sample handling degrades ctDNA before analysis begins
- Two persistent challenges: ctDNA is scarce in early-stage cancers, and clonal hematopoiesis (CHIP) can generate false-positive signals
What Is ctDNA Analysis?
ctDNA consists of short DNA fragments — typically fewer than 200 nucleotides, with peaks at 134 bp and 144 bp — shed by tumor cells into the bloodstream through apoptosis, necrosis, or active secretion. It's a tumor-specific subset of total cell-free DNA (cfDNA), which also contains DNA from healthy cells. Detecting ctDNA requires identifying tumor hallmarks: somatic mutations, aberrant methylation profiles, or characteristic fragmentation patterns.
Where ctDNA Fits in the Cancer Monitoring Landscape
ctDNA complements rather than replaces existing modalities:
- Tissue biopsies capture spatial tumor heterogeneity but are invasive, static, and sample only one tumor site
- Imaging (CT, MRI) measures anatomical changes but lacks molecular resolution and cannot detect MRD
- ctDNA provides systemic, dynamic, molecularly-specific data reflecting all disease sites in real time
Because ctDNA is shed from all tumor sites, it captures clonal diversity that a single-site biopsy may miss — a limitation that becomes critical in metastatic disease.
Detection Approaches
| Method | Sensitivity (VAF) | Use Case |
|---|---|---|
| Targeted PCR (qPCR, digital PCR, BEAMing) | 0.01–0.1% | Known mutations, rapid turnaround, MRD tracking |
| NGS panels (CAPP-Seq, targeted panels) | 0.004–1% | Comprehensive profiling, resistance detection |
| Methylation/fragmentomics | Variable | Tumor-agnostic screening, tissue-of-origin |
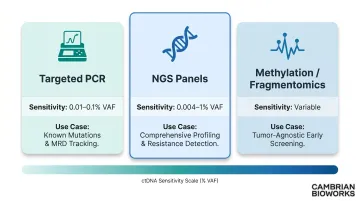

Each approach balances sensitivity, cost, breadth, and turnaround time. Targeted methods excel when specific mutations are known from prior profiling. NGS-based methods are better suited for tumor-agnostic settings or when comprehensive resistance profiling is needed.
Why ctDNA Monitoring Is Critical in Precision Oncology
Traditional RECIST imaging creates decision-making blind spots. It cannot detect MRD after surgery, cannot identify molecular progression before anatomical changes, and cannot capture emerging resistance mutations in real time. The median time from blood draw to RECIST progression in advanced genitourinary malignancies was 13.6 weeks — more than three months of potential treatment delay.
Real-Time Treatment Response Confirmation
Declining ctDNA levels during therapy indicate tumor regression, often earlier than imaging can confirm. In the phase III FLAURA trial, 68% of osimertinib-treated patients achieved ctDNA clearance by week 3 versus 44% on standard EGFR-TKIs, and clearance at week 3 predicted significantly longer progression-free survival.
MRD Detection Post-Surgery
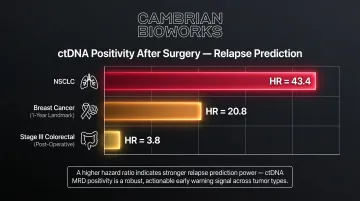
ctDNA positivity after curative-intent treatment predicts relapse with hazard ratios:
- Lung cancer (NSCLC): HR = 43.4 (p < 0.001)
- Breast cancer (1-year landmark): HR = 20.8 (95% CI 7.3-58.9)
- Stage III colorectal cancer (post-operative): HR = 3.8 (95% CI 2.4-21.0)
Hazard ratios at this magnitude mean ctDNA MRD status — positive or negative — can directly determine whether a patient receives adjuvant therapy after surgery.
On-treatment monitoring tells a different story. While MRD catches residual disease post-surgery, serial ctDNA testing during active therapy can detect acquired resistance before the tumor becomes clinically visible.
Resistance Mutation Surveillance
Emerging mutations appear in ctDNA before clinical progression:
- EGFR T790M in NSCLC: Detected a median of 2.2 months before progression
- ESR1 mutations in metastatic breast cancer: Detectable 6.7 months before clinical progression
A 6.7-month window is clinically meaningful — enough time to initiate a next-line agent, enroll in a trial, or reassess the treatment regimen before the patient's condition deteriorates.
Regulatory and Guideline Recognition
The FDA issued draft guidance in November 2024 on using ctDNA as a biomarker in early-stage solid tumor drug development. Multiple liquid biopsy companion diagnostics are FDA-approved, including cobas EGFR v2, FoundationOne Liquid CDx, and Guardant360 CDx.
ASCO's 2023 rapid recommendation update explicitly states "blood-based ctDNA is preferred owing to greater sensitivity" for ESR1 mutation testing in ER+/HER2- metastatic breast cancer — a strong recommendation backed by high-quality evidence.
How ctDNA Analysis Works – Step by Step
ctDNA analysis is not a single test but a multi-stage workflow. Errors at any step — especially pre-analytical steps — compromise downstream data and clinical interpretation.
Step 1: Collect the Blood Sample
Liquid biopsy typically requires 8–10 mL of peripheral blood in specialised cell-free DNA preservation tubes (e.g., Streck tubes, Roche BCT). Standard EDTA tubes must be processed within 4–6 hours to prevent white blood cell lysis; delayed processing to day 7 showed a 118-fold increase in genomic DNA contamination compared to preservation tubes.
Key quality metrics:
- Tube type compliance
- Temperature control during transport
- Processing timeline adherence
Collection timing relative to treatment matters: baseline samples before treatment, then serial samples during and after.
Step 2: Extract and Prepare cfDNA
Plasma is separated by centrifugation and cfDNA is purified using protocols optimised for short fragments (150–200 bp). At this stage, cfDNA yield, gDNA contamination levels, and fragment integrity directly determine whether downstream analysis is interpretable.
Insufficient cfDNA input is the leading technical cause of false-negative ctDNA results — particularly in early-stage cancers where tumour shedding is low. Tumour-derived fragments can be as short as 90 bp, so extraction kits optimised for the 166 bp nucleosome footprint may miss the shortest, most diagnostically relevant fragments.
Step 3: Select and Apply the Detection Method
Targeted PCR-based methods (qPCR, digital PCR, BEAMing):
- Best when specific mutations are known from prior profiling
- Sensitivity down to 0.01–0.1% variant allele frequency (VAF)
- Fast turnaround, cost-effective for serial monitoring
NGS panel-based approaches (CAPP-Seq, targeted panels):
- Better for tumour-agnostic settings or comprehensive resistance profiling
- Require higher cfDNA input and more complex bioinformatics
- Provide broader mutation landscape data
Non-mutation-based methods — methylation profiling and fragmentomics — are emerging complements that help overcome CHIP confounding (clonal haematopoiesis of indeterminate potential, where age-related blood cell mutations mimic tumour-derived signals).
Step 4: Analyse and Quantify ctDNA
Molecular response is assessed by:
- Tracking variant allele frequency (VAF) or tumour fraction over serial time points
- Evaluating ctDNA clearance (undetectable ctDNA post-treatment)
- Identifying emergence of new resistance mutations
- Comparing findings against baseline
The short half-life of cfDNA in circulation — estimated at 16 minutes to a few hours — means ctDNA levels reflect real-time tumour dynamics. This is a key advantage over imaging, which reflects accumulated anatomical change.
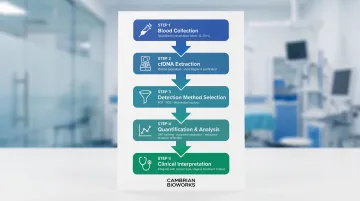
Step 5: Interpret Results and Inform Clinical Decisions
Interpreting ctDNA requires integrating molecular data with clinical context: tumour type, stage, treatment regimen, and timepoint.
Key interpretive scenarios:
- ctDNA clearance → response confirmation or MRD negativity
- Persistent or rising ctDNA → inadequate response or early relapse
- New mutation emergence → acquired resistance, may trigger treatment change
Critical pitfall: Over-interpreting a single ctDNA data point. A single undetectable result does not confirm remission; equally, a transient spike may not signal relapse. Clinical decisions should rest on consistent directional trends across at least two to three serial time points.
ctDNA Monitoring in Action: A Practical Scenario
Consider a stage IIIB NSCLC patient harbouring an EGFR exon 19 deletion, treated with first-line osimertinib. The oncologist uses serial ctDNA monitoring to guide treatment decisions at each stage.
| Timepoint | ctDNA Finding | Clinical Implication |
|---|---|---|
| Baseline (pre-treatment) | EGFR exon 19 deletion confirmed at detectable VAF | Establishes a trackable molecular marker for all future measurements |
| Week 4 (on-treatment) | Substantial drop or undetectable | Early molecular response; predicts treatment efficacy before imaging changes |
| Post-treatment surveillance | Remains undetectable | Consistent with MRD negativity and favourable prognosis |
| 6 months later | Re-emergence with EGFR T790M resistance mutation | Detected months before imaging progression; triggers proactive therapy switch |
The Week 4 finding reflects a pattern validated by clinical data. In the SWOG S1403 trial, complete plasma clearance at 8 weeks predicted PFS (HR 0.23, p < 0.0001), while RECIST response at the same timepoint was not predictive (HR 0.97, p = 0.90).
Each decision point in this scenario depends on data the lab generates. Reliable detection of low-abundance ctDNA requires extraction integrity, validated method sensitivity, and standardised pre-analytical protocols. Without these, the oncologist receives noise rather than actionable signal.
How Cambrian Bioworks Supports Reliable ctDNA Workflows
The most consequential pre-analytical variable in ctDNA analysis is cfDNA extraction quality. Cambrian Bioworks' Manta automated cfDNA extraction system addresses this gap in clinical and diagnostic labs.
Manta is CE-IVD marked (under Regulation EU 2017/746 IVDR) and ISO 13485 certified for quality management in medical device design and manufacturing. It completes cfDNA extraction in approximately 55 minutes with minimal genomic DNA contamination — critical for liquid biopsy applications where gDNA contamination can mask tumor-derived cfDNA signals.
Practical Workflow Benefits:
- Fits inside a biosafety hood at 43.5 cm × 28.2 cm × 26.2 cm — no dedicated instrument space required
- Pre-filled cartridges reduce hands-on time and simplify operator training
- Magnetic bead-based technology optimised for short-fragment cfDNA purification (150–200 bp)
- Processes 1 to 32 samples per run with no reagent waste from batching constraints
- Remote monitoring capability supports multi-instrument oversight and operational efficiency
- Integrated UVC lamp provides in-system contamination control
These features reduce the technical variability that remains a key barrier to routine ctDNA adoption. Standardising the extraction step means downstream results are more reproducible across runs and operators — and the performance data backs this up.
Validated Performance:
In comparative studies across seven clinical patient samples, Manta's OncoSeek cfDNA extraction kit delivered 5–20× higher cfDNA yield using just 1 mL of plasma versus a leading competitor requiring 2 mL. The cell-free fraction (100–200 bp) exceeded 45% in tumour-positive samples, confirmed by TapeStation cfDNA ScreenTape analysis.

Manta-extracted cfDNA is validated for use with NGS, qPCR, and digital PCR, covering the primary downstream platforms for mutation profiling, MRD detection, and resistance surveillance.
Scaling ctDNA Testing:
As labs scale up ctDNA testing volume, the combination of Manta for cfDNA extraction and Beluga for automated liquid handling supports higher throughput without sacrificing per-sample quality. Both systems handle 1–32 samples per run, so labs can scale incrementally without retooling their existing workflow.
Trusted by 67+ labs globally — including Tata Memorial Hospital and ACTREC — Manta gives oncology and diagnostic labs a validated, reproducible starting point for every ctDNA test they run.
Frequently Asked Questions
What is the difference between circulating tumor DNA and cell-free DNA?
Cell-free DNA (cfDNA) is the broader category of fragmented DNA circulating in blood, shed by all types of cells through normal apoptosis. ctDNA is specifically the tumor-derived fraction of cfDNA, carrying tumor-specific genetic alterations. In cancer patients, ctDNA can range from less than 1% of total cfDNA in early-stage disease to over 90% in advanced metastatic cases.
How is ctDNA detected?
ctDNA is detected from blood plasma after cfDNA extraction, using either targeted methods (digital PCR, qPCR) that search for known tumor mutations with very high sensitivity, or NGS-based approaches that profile a broader range of genomic alterations. Newer methods also detect ctDNA through DNA methylation patterns and fragmentation profiles, which don't require prior knowledge of tumor mutations.
How do you interpret ctDNA results?
ctDNA results are most meaningful when interpreted longitudinally — tracking trends across serial time points rather than relying on a single value. Declining or undetectable ctDNA generally indicates response or MRD negativity; rising ctDNA or new mutations may signal inadequate response, early relapse, or acquired resistance, with clinical context (tumor type, stage, treatment regimen, and timing) essential for accurate interpretation.
What does a positive ctDNA test mean?
A positive ctDNA result means tumor-derived DNA has been detected in the blood, indicating active disease — whether from an actively shedding tumor, residual disease after treatment (MRD positivity), or recurrence. Any positive finding warrants clinical follow-up and should be interpreted alongside imaging and patient history.
How accurate is the ctDNA test?
ctDNA test accuracy varies by cancer type, stage, detection method, and tumour shedding rates. Sensitivity is markedly higher in advanced disease — approximately 82% in metastatic cancers versus 47% in localised tumours. Specificity challenges from CHIP variants (non-tumour mutations from ageing blood cells) are mitigated by rigorous pre-analytical preparation and error-correction sequencing.
What is considered a high ctDNA level?
ctDNA levels are typically expressed as variant allele frequency (VAF) or tumour fraction. Levels above 10% tumour fraction are often considered elevated and have been linked to poorer prognosis, though "high" is cancer-type specific and always interpreted relative to that patient's own baseline — universal reference ranges do not yet exist.


