Introduction
Oncologists face a persistent challenge: cancer is not a single disease but a collection of heterogeneous, evolving molecular landscapes. Tumors mutate under therapeutic pressure, shedding clones that resist first-line treatments. Traditional tissue biopsies capture only a single snapshot from one tumor region at one point in time, leaving clinicians blind to the genetic diversity lurking across metastatic sites and the dynamic shifts occurring during therapy. This static molecular portrait complicates treatment decisions — particularly when archival tissue is years old, unavailable, or insufficient for comprehensive genomic profiling.
Plasma circulating tumor DNA (ctDNA) offers a minimally invasive solution. Detectable in a simple blood draw, ctDNA provides a real-time, whole-tumor genomic view by integrating signals from multiple tumor sites simultaneously.
This blog covers the biology of ctDNA, the analytical workflow from blood draw to clinical insight, and key applications in precision oncology — including early detection, therapy selection, treatment monitoring, and minimal residual disease surveillance — along with the challenges and emerging directions defining the field.
TLDR
- ctDNA is a tumor-derived subset of cfDNA detectable in plasma, carrying cancer-specific mutations
- Enables early detection, therapy selection, treatment response monitoring, and minimal residual disease surveillance without invasive biopsies
- Captures tumor heterogeneity better than single-site tissue biopsies by sampling DNA from multiple tumor foci
- Pre-analytical variables directly affect ctDNA test reliability, making high-quality cfDNA extraction non-negotiable
What Is Plasma ctDNA and How Does It Differ from cfDNA?
Cell-Free DNA: The Foundation
Cell-free DNA (cfDNA) is fragmented DNA circulating in blood plasma, released by normal cell turnover, apoptosis, and necrosis. These fragments average approximately 166 base pairs in length, reflecting the nucleosome-protected DNA footprint left behind during programmed cell death. cfDNA is present in all individuals, healthy or diseased, and has a short half-life—approximately 16 minutes to 2.5 hours—making it a real-time snapshot of physiological and pathological states.
Circulating Tumor DNA: The Tumor Signal
ctDNA is a subset of cfDNA originating specifically from tumor cells—whether from the primary tumor, metastatic lesions, or circulating tumor cells. Its defining characteristics include:
- Carries tumor-specific somatic mutations, copy number alterations, and methylation changes
- Fragments typically shorter than non-mutant cfDNA, enriched in the 150–200 bp range — aiding selective recovery during extraction
- Represents less than 1% of total cfDNA in early-stage cancers, rising with disease burden
- Correlates with tumor volume and stage in metastatic patients
Despite this correlation, approximately 15% of metastatic patients still have undetectable ctDNA — a finding often associated with better overall survival.
Plasma vs. Serum: Why the Matrix Matters
Plasma is the preferred matrix for ctDNA testing because serum processing involves clotting, which can lyse white blood cells and release high-molecular-weight genomic DNA. This contamination dilutes the ctDNA signal, reducing sensitivity and specificity. The blood collection and processing protocol directly impacts test quality — which is why standardised pre-analytical workflows are non-negotiable in clinical ctDNA testing.
The ctDNA Workflow: From Blood Draw to Clinical Insight
Pre-Analytical Phase: Blood Collection and Processing
Blood is typically collected in EDTA or Streck Cell-Free DNA BCT tubes, which stabilize white blood cells and prevent genomic DNA release. Delays in processing or use of improper tube types are among the most common sources of genomic DNA contamination, degrading ctDNA signal quality. After collection, blood undergoes centrifugation to separate plasma from cellular components, followed by immediate storage at –80°C to preserve cfDNA integrity.
cfDNA Extraction: Selective Recovery of Short Fragments
Magnetic bead-based isolation selectively recovers short cfDNA fragments from plasma while minimizing contamination from high-molecular-weight genomic DNA released by white blood cells. Extraction yield, purity, and fragment-size profiles directly determine downstream assay performance.
Cambrian Bioworks' Manta, for instance, automates this step end-to-end for clinical labs. Key design features include:
- CE-IVD certified for clinical use, completing extraction in ~27–30 minutes
- Pre-filled cartridges that standardize the workflow and reduce operator error
- Selective binding conditions that preferentially capture 150–200 bp cfDNA fragments
- Output compatible with sensitive oncology applications, including liquid biopsy and minimal residual disease detection

Clean, high-quality cfDNA at this stage is the foundation for everything that follows in the workflow.
Library Preparation and Sequencing
Extracted cfDNA is converted into sequencing libraries and subjected to targeted panel sequencing—often covering cancer driver genes—or whole-exome sequencing at high depth. Targeted panels commonly run at 1,000× coverage or more. Unique molecular identifiers (UMIs) tag individual DNA molecules before amplification, distinguishing true mutations from sequencing artifacts—a critical safeguard when detecting low-frequency somatic variants below 1% allele frequency.
Bioinformatics and Reporting
Variant calling is performed against germline DNA (typically buffy coat) to filter out non-tumor variants. Bioinformatics pipelines estimate ctDNA fraction (tumor fraction), calculate tumor mutational burden (TMB), and translate findings into a clinical report that guides treatment decisions.
Key Clinical Applications of Plasma ctDNA in Precision Medicine
Early Cancer Detection and Screening
ctDNA can harbor cancer-associated mutations detectable in plasma before clinical symptoms arise. Research has shown mutations detectable in blood up to two years before cancer diagnosis, enabling multi-cancer early detection (MCED) tests. These tests interrogate ctDNA methylation patterns and fragmentation profiles to identify cancer type and tissue of origin, even when the primary tumor site is unknown. MCED platforms like Galleri analyze signals across 50+ cancer types, offering the potential for population-level screening in asymptomatic individuals.
Therapy Selection and Biomarker-Driven Treatment
ctDNA profiling reveals actionable somatic alterations that guide targeted therapy selection. In metastatic bladder cancer, ctDNA reproduces the driver gene landscape seen in tissue biopsies with approximately 83% concordance — making it a viable companion diagnostic when tissue is unavailable or insufficient.
Key actionable mutations identified through ctDNA profiling include:
- FGFR3 and ERBB2 — matched targeted therapies
- EGFR and KRAS — guides immunotherapy or targeted agent selection
- ERCC2 — associated with platinum sensitivity in urothelial cancers
Tumor mutational burden (TMB) estimation from ctDNA is also gaining ground as an immunotherapy biomarker. ctDNA-derived TMB correlates well with tissue TMB, providing a non-invasive option for patient stratification in immunotherapy trials.
Treatment Response Monitoring
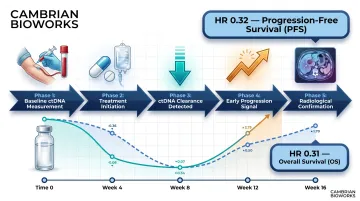
Serial ctDNA measurements during therapy give oncologists an early, quantitative readout of tumor response. Levels drop with effective treatment and rise before imaging detects progression — providing a lead over conventional radiology. A meta-analysis of 32 studies involving 3,047 non-small cell lung cancer patients demonstrated that ctDNA clearance during treatment was associated with a hazard ratio of 0.32 for progression-free survival and 0.31 for overall survival, confirming ctDNA as a powerful prognostic tool.
Minimal Residual Disease Detection and Relapse Surveillance
Minimal residual disease (MRD) refers to ctDNA detectable in blood after curative-intent treatment — surgery, radiation, or neoadjuvant chemotherapy. Post-treatment ctDNA positivity indicates residual disease or micro-metastases not visible on imaging and is associated with increased relapse risk. In colon cancer, ctDNA positivity after resection predicts recurrence-free survival outcomes, enabling earlier intervention or adjuvant therapy decisions.
Monitoring Clonal Evolution and Acquired Resistance
Resistance mutations rarely announce themselves — but ctDNA catches them early. Serial profiling over the treatment course can capture acquired resistance before clinical progression becomes apparent. Monitoring EGFR T790M mutations in NSCLC patients showed a median lead time of 1.1 months before radiological progression, with some patients identified up to 16.6 months earlier.
Unlike a single tissue biopsy, ctDNA integrates DNA shed from multiple tumor foci simultaneously — capturing the full landscape of clonal evolution. Studies show 90% of driver mutations are consistently detected across serial ctDNA samples, compared to significantly lower concordance in serial tissue samples.
ctDNA vs. Tumor Tissue Biopsy: A Practical Comparison
Accessibility: Static vs. Dynamic Molecular Portrait
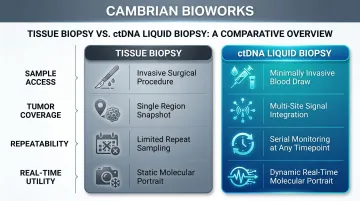
Tissue biopsy captures a static snapshot — one region, one timepoint, with no ability to track how a tumor changes under treatment pressure. ctDNA liquid biopsy, by contrast, provides a dynamic molecular portrait that evolves with the patient.
Key practical differences include:
- Sample access: Blood draw vs. surgical or invasive procedure
- Tumor coverage: Integrates signal from multiple metastatic sites vs. single biopsy region
- Repeatability: Serial monitoring at any timepoint vs. limited repeat sampling
- Real-time utility: Tracks treatment response and clonal evolution as it happens
Concordance and Complementarity
Mutation concordance between ctDNA and matched tumor tissue is high, approximately 83% in published cohorts for driver gene alterations. However, discordances do exist. Subclonal mutations and intra-tumor heterogeneity mean that ctDNA occasionally detects variants absent from archival primary tissue and vice versa. A complementary approach—where ctDNA is primary and tissue biopsy serves as fallback when ctDNA levels are undetectable—is recommended.
When Tissue Biopsy Remains Essential
That complementary framework has clear boundaries. When ctDNA levels are very low or undetectable, or when RNA-based biomarkers — tumor subtyping, PD-L1 expression — are needed for treatment decisions, tissue biopsy remains essential. Low ctDNA in metastatic disease is itself associated with better overall survival, suggesting less aggressive biology.
Challenges and Limitations of ctDNA Testing
The Low Abundance Problem
ctDNA can represent less than 1% of total cfDNA in early-stage or treated patients. This makes false negatives possible and requires deep sequencing—often 1,000× or more—to achieve sufficient sensitivity. Ultra-sensitive detection methods like ddPCR and error-corrected NGS are essential, as is high-quality input DNA. Typical ctDNA fractions range from <0.1% in early-stage disease to >10% in metastatic cancers.
Clonal Hematopoiesis Confound
Somatic mutations arising in aging hematopoietic stem cells (clonal hematopoiesis of indeterminate potential, or CHIP) can mimic ctDNA mutations in plasma, generating false-positive calls. Studies show that up to 17% of mutations detected in plasma may be CHIP-related, affecting 29% of patients, with rates reaching 20% in individuals over age 70.
Labs mitigate this by sequencing matched white blood cell (buffy coat) DNA as a germline reference and requiring tumor-specific variant allele frequency (VAF) thresholds above background. Rigorous pre-analytical workflows and careful plasma-cellular fraction separation are therefore non-negotiable — a point where extraction quality directly determines result reliability. Platforms with minimal genomic DNA contamination, such as Cambrian Bioworks' Manta, reduce this interference at the source.
Standardization Gaps
ctDNA testing still lacks consensus across several dimensions:
- No universal pre-analytical standards for sample collection or processing
- No approved clinical decision-making cutoffs in most cancer types outside select FDA/CE-IVD assays
- Inconsistent concordance across commercial platforms for low-VAF variants
Ongoing regulatory and guideline-setting efforts by ESMO, ASCO, and the Association for Molecular Pathology are working to address these gaps.
The Future of Plasma ctDNA in Precision Oncology
Emerging Directions: Methylation, Fragmentomics, and Multi-Modal Biomarkers
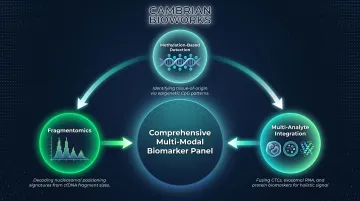
Three technological shifts are converging to push ctDNA beyond single-analyte testing toward comprehensive multi-modal biomarker panels:
- Methylation-based detection: Multi-cancer early detection tests that identify tissue-of-origin through tumor-specific methylation signatures
- Fragmentomics: Using cfDNA fragment length patterns and nucleosome positioning to infer cancer type without relying on mutations alone
- Multi-analyte integration: Combining ctDNA with circulating tumor cells, exosomes, and protein markers for higher sensitivity and specificity
This convergence is reflected in market momentum. The global liquid biopsy market was valued at $3.65 billion in 2024 and is projected to reach $7.05 billion by 2030, growing at a CAGR of 11.8%.
Routine Clinical Integration and Global Adoption
Falling sequencing costs, standardized extraction workflows, and expanding regulatory approvals are collectively moving ctDNA testing out of research settings and into routine oncology practice. The pace is especially pronounced in Asia. In the Asia-Pacific region, the liquid biopsy market is projected to reach $4.57 billion by 2033, growing at a CAGR of 31.23%.
Labs in India and other emerging markets are increasingly adopting liquid biopsy workflows. For these facilities, access to CE-IVD certified, affordable, and automated extraction tools — such as Cambrian Bioworks' Manta — directly determines whether high-quality ctDNA testing is operationally viable at scale.
Frequently Asked Questions
Is ctDNA in plasma?
Yes, ctDNA circulates in blood plasma as a subset of cell-free DNA (cfDNA). It is released by tumor cells through apoptosis, necrosis, and active secretion, and can be isolated from plasma by centrifuging whole blood to remove cellular components.
What does a positive ctDNA mean?
A positive ctDNA result means tumor-derived DNA carrying cancer-associated mutations was detected in plasma. This can indicate active cancer, residual disease after treatment, or early relapse — with clinical significance varying by whether the test was used for screening, treatment monitoring, or post-treatment surveillance.
What to do if ctDNA is positive?
A positive ctDNA result should prompt consultation with an oncologist to guide next steps based on stage and setting, which may include confirmatory imaging, tumor tissue biopsy, adjustment of treatment, or enrollment in a biomarker-driven clinical trial.
How much does a ctDNA blood test cost?
ctDNA test costs vary by country, clinical setting, and assay type. In India, panel-based liquid biopsy tests typically range from ₹15,000 to ₹80,000; in Europe costs are broadly comparable or lower. US pricing runs higher, often between $1,000 and $5,000, though costs across all markets are falling as sequencing throughput improves.
How is ctDNA different from cfDNA?
cfDNA is all fragmented DNA circulating in blood from any cell type, while ctDNA is specifically the tumor-derived fraction of cfDNA that carries cancer-specific somatic mutations. ctDNA is a subset of cfDNA and typically represents a small and variable proportion of that pool.
What cancers can be detected by ctDNA testing?
ctDNA testing has been studied across many cancer types, including lung, colorectal, breast, bladder, prostate, and ovarian cancers. ctDNA detectability varies by cancer type, stage, and tumor burden, with metastatic and high-stage cancers typically shedding more detectable ctDNA than early-stage disease.


