Introduction
Cell-free tumor DNA (ctDNA) in blood plasma is now a frontline tool in oncology diagnostics. A single blood draw can enable non-invasive tumor profiling, real-time treatment monitoring, and early recurrence detection — capabilities that are changing how laboratories approach cancer diagnostics at every stage of care.
Yet the signal only has value when interpreted correctly. A trace mutation at 0.5% variant allele frequency (VAF) could indicate residual disease requiring immediate intervention — or it could be a false positive from clonal hematopoiesis. Similarly, a spike in total cfDNA concentration may reflect metastatic progression, or nothing more than pre-draw exercise.
For laboratory professionals, correct interpretation depends on a clear grasp of ctDNA biology, expected concentration ranges across disease states, and the analytical thresholds that define assay performance. Pre-analytical variables can degrade results without obvious flags. When they go unrecognised — or when total cfDNA is conflated with ctDNA, VAF detection limits are ignored, or clonal hematopoiesis is overlooked — the clinical consequences are direct.
This guide covers:
- The biology of ctDNA and how it enters circulation
- Concentration ranges across healthy and disease states
- Key physical and molecular properties governing behaviour
- Measurement considerations and assay performance thresholds
- Practical implications of aberrant ctDNA signals
TL;DR
- ctDNA is tumor-derived cfDNA carrying cancer-specific mutations — circulating at concentrations as low as <0.1% VAF in early-stage disease
- Healthy adults have <10 ng/mL total cfDNA; ctDNA fraction ranges from <0.1% (early-stage) to >10% (metastatic)
- Levels fluctuate with tumor burden, treatment response, clonal hematopoiesis, and pre-analytical handling errors
- Reliable detection requires matched assay sensitivity (0.01–5% VAF by platform) and rigorous extraction workflows to prevent genomic contamination
- Ignoring CHIP, applying wrong VAF thresholds, or relying on a single time-point creates both false positives and false negatives
What ctDNA Represents in Blood Plasma
Technical Definition and Molecular Characteristics
Understanding ctDNA starts with distinguishing it from the broader pool of cell-free DNA circulating in plasma.
ctDNA is the tumor-derived fraction of total cfDNA, defined by cancer-specific molecular alterations. It originates primarily from apoptotic or necrotic tumor cells — and in smaller quantities from tumor-derived exosomes — carrying somatic mutations, copy number alterations, methylation changes, and chromosomal rearrangements unique to the tumor.
Key molecular features:
- Somatic mutations: show up to 90% concordance with matched solid tumors
- Copy number alterations: detectable, though clonal hematopoiesis complicates interpretation
- Cancer-related hypo/hypermethylation patterns that serve as tissue-of-origin markers
- Chromosomal rearrangements, including chromosomal instability and extrachromosomal DNA
The cfDNA-ctDNA Relationship
All ctDNA is cfDNA, but only a small fraction of total cfDNA is tumor-derived. In healthy individuals, approximately 76–79% of cfDNA originates from leukocytes, with minor contributions from organ turnover and inflammation.
This hematopoietic dominance persists even when total cfDNA concentration exceeds normal levels by 10-fold. The result is a fundamental specificity challenge: isolating the tumor signal from overwhelming background noise.
Clinical Applications
| Application | Purpose | Key Assays / Evidence |
|---|---|---|
| Companion diagnostics | Mutation profiling for targeted therapy selection | cobas EGFR Mutation Test v2 (FDA-approved 2016); Guardant360 CDx; FoundationOne Liquid CDx |
| Minimal residual disease monitoring | Post-surgical detection of residual disease to guide adjuvant therapy | Signatera (CRC): sensitivity >65%, specificity 99.9% |
| Resistance detection | Real-time tracking of emerging resistance mutations during treatment | EGFR T790M in NSCLC; ERBB2, TP53, PIK3CA alterations |
Factors That Influence ctDNA Levels in Real-World Operation
Tumor Location and Shedding Biology
ctDNA concentration does not simply mirror tumor size. Bettegowda et al. demonstrated that in advanced disease, ctDNA is detectable in >75% of patients with pancreatic, ovarian, colorectal, bladder, gastroesophageal, breast, melanoma, and hepatocellular cancers. Primary brain tumors (gliomas), however, show ctDNA detection in <10% of peripheral blood samples — a direct consequence of the blood-brain barrier limiting systemic shedding.
Key modulators of shedding include:
- Highly vascular tumors release more ctDNA into circulation
- CNS tumors behind the blood-brain barrier shed at minimal levels
- Apoptotic and necrotic cell turnover drives the bulk of ctDNA release
- Therapy-induced cell death causes a temporary spike in ctDNA, then decline
Pre-Analytical Variables
Blood collection tube type and processing delays exert direct, measurable effects on ctDNA signal integrity.
Tube type stability:
- K2EDTA tubes — stable up to 6 hours at room temperature; gDNA contamination increases significantly after 24 hours
- Streck cfDNA BCT — maintains cfDNA stability for at least 3 days at room temperature, with manufacturer claims extending to 14 days at 6-37°C
Processing delay effects (K3EDTA tubes at room temperature):
- Total cfDNA concentration increases gradually due to leukocyte lysis
- Mutant allele fraction (MAF) decreases as wild-type gDNA dilutes tumor signal
- Absolute ctDNA concentration (copies/mL) remains relatively stable
- Significant cfDNA elevation occurs after 48 hours
Beyond handling artifacts, a biological variable presents an equally serious challenge for accurate ctDNA interpretation.
Clonal Hematopoiesis: The Critical Biological Confounder
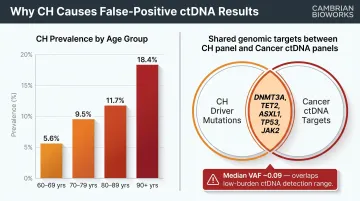
Clonal hematopoiesis (CH) prevalence rises sharply with age, representing one of the primary sources of false-positive ctDNA signals:
| Age Group | CH Prevalence |
|---|---|
| 60-69 years | 5.6% (95% CI: 5.0-6.3) |
| 70-79 years | 9.5% (95% CI: 8.4-10.8) |
| 80-89 years | 11.7% (95% CI: 8.6-15.7) |
| 90+ years | 18.4% (95% CI: 12.1-27.0) |
This matters clinically because CH-associated mutations create signals that are nearly indistinguishable from low-level tumor DNA:
- The most frequently mutated genes — DNMT3A, TET2, ASXL1, TP53, JAK2 — overlap directly with standard cancer mutation panels
- Median VAF of CH mutations sits at 0.09, the same range where low-burden ctDNA carries the most clinical value
- Without matched white blood cell sequencing, hematopoietic-origin mutations are routinely misclassified as tumor-derived
Normal vs. Elevated ctDNA Levels: Understanding the Diagnostic Range
ctDNA operates across a wide dynamic range depending on disease state, cancer type, and tumor burden. There is no universal threshold; interpretation requires context-specific reference ranges.
Baseline cfDNA Levels in Healthy Individuals
Total plasma cfDNA in healthy adults typically measures below 10 ng/mL:
- Healthy cohort 1 (N=812): Mean 4.3 ± 8.6 ng/mL (median 2.9 ng/mL)
- Healthy cohort 2 (N=64): Mean 6.0 ± 10.5 ng/mL (median 3.4 ng/mL)
At this baseline, true ctDNA is generally undetectable or present at sub-0.1% VAF with standard assays. However, cfDNA levels increase physiologically with age, exercise, and inflammation, and pathologically in myocardial infarction, stroke, and autoimmune conditions.
Exercise-induced spikes: Intensive exercise elevates cfDNA 5- to 39-fold above baseline, primarily from neutrophil turnover, returning to baseline within 1 hour. Blood draw timing relative to physical activity represents a potential confounder requiring standardization.
Tumor-Derived ctDNA Across Disease Stages
Stage-stratified detection rates from Bettegowda et al. across multiple cancer types:
| Stage | Detection Rate | Mutant Fragments (per 5 mL plasma) |
|---|---|---|
| Stage I | 47% | 10¹–10² |
| Stage II | 55% | ~10² |
| Stage III | 69% | ~10³ |
| Stage IV | 82% | 10⁴–10⁵ |
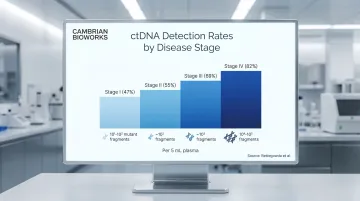
ctDNA fraction within total cfDNA follows a parallel trajectory: typically <1% in early-stage solid tumors, rising to several percent in locally advanced disease, and potentially exceeding 10-40% in metastatic settings.
Safe Operating Margin for Clinical Decision-Making
Assays carry platform-specific limits of detection (LOD), and reporting variants at or near these thresholds produces clinically misleading results. Detection sensitivity varies substantially by platform:
- Standard NGS panels: 1–2% VAF
- Digital PCR: 0.001–0.1% VAF
- Error-corrected platforms (e.g., iDES-enhanced CAPP-Seq): down to 0.00025% VAF
When ctDNA VAF falls below a given assay's sensitivity floor, false positives increase — driven by sequencing artifacts, CHIP-derived variants, or poor sample quality. Concordance between assays drops sharply below 1% VAF. ctDNA-to-tissue concordance ranges from 60–80% overall, with notably poorer consistency when mutant allele fraction is below 1%.
Key Biological Properties That Define ctDNA as a Biomarker
ctDNA's clinical utility is governed by four interconnected biological properties that determine how it can and cannot be used.
Property 1: Fragment Size and Nucleosomal Footprint
ctDNA fragments are typically shorter (~145-167 bp) than germline cfDNA, reflecting apoptotic cleavage patterns. Tumor-derived fragments are often even shorter, which can be exploited for enrichment but also means they are prone to degradation during extraction.
The peak cfDNA fragment size is 167 bp, corresponding to DNA wrapped around one nucleosome (147 bp) plus linker DNA (~20 bp). Targeting fragments shorter than 145 bp can improve ctDNA detection sensitivity, achieving up to 2-fold median ctDNA enrichment by analyzing the 90-150 bp size range.
Property 2: Short Circulatory Half-Life
cfDNA has a half-life estimated at 5-150 minutes in circulation. This rapid clearance means ctDNA levels track current tumor activity closely, making it a sensitive indicator of treatment response. However, this also means sample handling delays introduce significant pre-analytical bias — processing delays beyond 6 hours in EDTA tubes at room temperature cause genomic DNA contamination that dilutes the tumor signal.
Property 3: Tumor Specificity and Clonal Heterogeneity
ctDNA carries mutations from multiple tumor clones simultaneously, reflecting spatial and temporal heterogeneity. This is both a strength (global tumor snapshot vs. single-site biopsy) and a complexity — interpreting multiple co-occurring variants requires clinical context.
Polyclonal shedding from metastatic sites captures the full tumor burden, but VAF values differ by location:
- Non-CNS metastases in highly vascular organs yield the highest VAF
- Brain metastases show lower VAF due to the blood-brain barrier limiting shedding into systemic circulation
How ctDNA is Measured, Extracted, and Validated
ctDNA analysis is a two-stage technical challenge. The first stage is extracting intact, high-yield cfDNA from plasma without genomic DNA contamination. The second is applying a sensitive sequencing or PCR-based method to detect low-frequency tumor variants.
Measurement Methods
Three detection approaches dominate clinical and research workflows:
- Targeted deep sequencing panels — for known mutation hotspots; sensitivity 0.02-2% VAF depending on error-correction
- Digital PCR — for quantification of specific variants; sensitivity 0.001-0.1% VAF
- Methylation-based assays — for tissue-of-origin calling; Liu et al. demonstrated simultaneous detection and localization of >50 cancer types
| Platform | Minimum Detectable VAF | Technology |
|---|---|---|
| Standard NGS panels | 1-2% | Amplicon/hybrid-capture |
| TAM-seq | 2% | Tagged amplicon deep sequencing |
| ddPCR | 0.001-0.1% | Droplet digital PCR |
| BEAMing | 0.01% | Bead emulsion PCR |
| CAPP-Seq | 0.02% | Error-corrected NGS |
| iDES-enhanced CAPP-Seq | 0.00025% | Integrated digital error suppression |
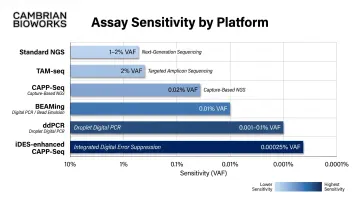
Method choice must match the clinical question: companion diagnostics typically require 1-2% sensitivity, while minimal residual disease monitoring demands detection below 0.1% VAF.
That sensitivity ceiling, however, is only achievable if extraction quality holds up upstream.
Extraction as a Rate-Limiting Step
Downstream assay performance is directly constrained by extraction quality. Insufficient yield, genomic DNA contamination, or fragment loss during purification lowers effective sensitivity, leaving even high-sensitivity sequencing platforms unable to detect low-burden ctDNA.
CE-IVD certified automated platforms like Manta by Cambrian Bioworks address this directly — designed to maximize cfDNA yield from plasma while minimizing genomic contamination for reproducible, clinical-grade liquid biopsy workflows.
Manta uses magnetic bead-based purification optimized for short cfDNA fragments (150-200 bp), completing extraction in approximately 55 minutes with minimal hands-on time.
Interpreting Aberrant ctDNA Signals: Causes and Consequences
The Differential Interpretation Challenge
Elevated ctDNA beyond expected range for a given disease state may indicate:
- Disease progression or metastatic expansion
- Treatment resistance or clonal evolution
- Clonal hematopoiesis (false positive)
- Inflammation or tissue injury (cfDNA elevation without tumour signal)
- Assay artifact from poor sample quality
Each signal requires interpretation against clinical context, longitudinal trends, and technical quality metrics — no single result is conclusive in isolation.
Performance Degradation at Low Tumour Fractions
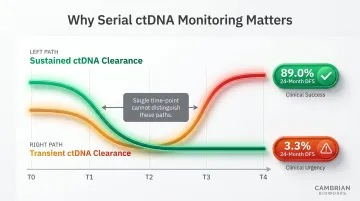
When ctDNA VAF falls below assay sensitivity thresholds — such as during post-treatment minimal residual disease monitoring — false negatives increase, leading to incorrect inference of complete remission. In the GALAXY study of 2,240 colorectal cancer patients, sustained ctDNA clearance after adjuvant chemotherapy yielded 89.0% 24-month disease-free survival vs. 3.3% for transient clearance, demonstrating that dynamics — not single time-points — drive prognostic value.
Clinical and Compliance Implications
In companion diagnostic contexts, ctDNA results directly guide therapy selection. The FDA approved the cobas EGFR Mutation Test v2 in 2016 for detecting EGFR mutations in NSCLC patients to guide TKI therapy, with Guardant360 CDx and FoundationOne Liquid CDx following as broader liquid biopsy panels.
A false negative at low tumour burden may deny a patient access to targeted therapy. Regulatory bodies require companion diagnostic ctDNA assays to meet defined LOD, analytical sensitivity, and clinical concordance thresholds before approval — making pre-analytical and extraction consistency critical upstream of the assay itself.
Common Misinterpretations of ctDNA in Practice
Treating Total cfDNA Elevation as Equivalent to ctDNA Elevation
Many labs and clinicians conflate high cfDNA concentration with confirmed tumor shedding, without accounting for benign sources that elevate cfDNA independently of tumor activity:
- Exercise and physical stress — can transiently spike cfDNA from muscle and immune cell turnover
- Systemic inflammation — infection, autoimmune flares, and tissue injury all contribute
- Clonal hematopoiesis — introduces somatic mutations that mimic tumor-derived variants
Specificity requires mutation confirmation, not just quantity. A patient recovering from myocardial infarction or a recent marathon may show cfDNA levels exceeding 100 ng/mL with no tumor-derived signal whatsoever.
Applying a Uniform Detection Threshold Across Assays and Cancer Types
A VAF cutoff validated for one platform or tumor type cannot be directly transferred to another. Ignoring assay-specific limits of detection and sequencing depth requirements is a primary driver of inter-lab discordance.
Concordance between liquid biopsy and tissue biopsy ranges widely — from 43% to over 86% positive predictive value — depending on the platform used, the gene targets interrogated, and the tumor fraction present in the sample.
Overlooking Temporal and Sampling Dynamics
A single ctDNA measurement is a snapshot, not a trajectory. The predictive and prognostic value of ctDNA derives from longitudinal trends — rising, falling, or emerging variant patterns. Clinical decisions based on a single time point without baseline context can be misleading.
The GALAXY study makes this concrete. Sustained clearance and transient clearance look identical at a single time point — yet their 24-month DFS outcomes diverge sharply: 89.0% vs. 3.3%. Spontaneous clearance without chemotherapy occurred in just 1.9% of MRD-positive patients. Serial monitoring reveals what any individual draw cannot.
Frequently Asked Questions
What is cfDNA in blood plasma?
Cell-free DNA (cfDNA) is fragmented DNA circulating in blood plasma, released by normal cellular apoptosis and necrosis. In healthy individuals, it originates primarily from hematopoietic cells, while in disease states it can include tumor-derived DNA (ctDNA).
Can cancer be detected in plasma?
Yes, cancer-derived DNA (ctDNA) can be detected in plasma through liquid biopsy using deep sequencing or digital PCR to identify tumor-specific mutations. Sensitivity depends on cancer type, stage, and tumor burden. Early-stage detection remains technically challenging due to low ctDNA fractions.
What is the normal range for cell-free DNA?
Total cfDNA in healthy adults is typically below 10 ng/mL of plasma. In cancer-free individuals, ctDNA is generally undetectable or below 0.1% VAF — a fraction that rises sharply with active malignancy.
What is the difference between cfDNA and ctDNA?
cfDNA is the broad category of all cell-free DNA in circulation, while ctDNA is a tumor-specific subset carrying somatic mutations. ctDNA represents a small and variable fraction of total cfDNA, especially in early-stage disease where it may constitute less than 1% of total circulating DNA.
What factors affect ctDNA levels in blood?
ctDNA levels are influenced by tumor burden, stage, vascularity, treatment response, clonal hematopoiesis, and pre-analytical variables — including blood tube type (EDTA vs. Streck), processing time, and extraction methodology. Exercise, inflammation, and tissue injury can elevate total cfDNA without raising the tumor-specific signal.
How is ctDNA used to monitor cancer treatment?
Serial ctDNA measurements track variant allele frequency over time. Declining levels suggest treatment response; persistent or rising levels indicate resistance or residual disease. Emerging new variants — such as EGFR T790M in NSCLC on TKI therapy — can flag acquired resistance before clinical progression.


