Introduction
Clinical laboratories face a fundamental challenge in liquid biopsy workflows: extracting enough high-quality cell-free DNA (cfDNA) from plasma samples to detect mutations present in less than 1% of total DNA. This challenge is compounded by cfDNA's inherently low concentration — healthy individuals typically have just 1-10 ng/mL of cfDNA in plasma, while early-stage cancer patients may have only marginally higher levels. Even small extraction losses can render samples unsuitable for mutation detection.
This guide is for molecular diagnostics labs, clinical researchers, oncology facilities, prenatal testing centres, and transplant medicine centres that depend on cfDNA as a liquid biopsy biomarker. Most labs focus on optimising downstream analysis, yet extraction quality is the variable most likely to determine whether an assay succeeds or fails — and it remains consistently underestimated.
Poor extraction directly causes false negatives in minimal residual disease monitoring, inflates no-call rates in non-invasive prenatal testing (NIPT), and compromises circulating tumour DNA (ctDNA) detection sensitivity.
What follows is a practical, end-to-end breakdown of cfDNA extraction: the variables that determine quality, the steps where labs most often lose yield, and what separates a reliable workflow from one that generates inconclusive results.
TL;DR
- cfDNA extraction isolates short (~167 bp) DNA fragments used for cancer detection, prenatal testing, and transplant monitoring
- Plasma is the gold standard matrix — serum causes significant genomic DNA contamination due to clotting-induced cell lysis
- Silica membrane kits offer higher total yield; magnetic bead-based systems deliver better ctDNA enrichment and automation compatibility
- Automated extraction reduces inter-operator variability and is essential for clinical-grade workflows
- Key quality parameters include total yield, genomic DNA contamination, and fragment size integrity
What Is cfDNA Extraction from Plasma?
cfDNA extraction from plasma is the targeted isolation of cell-free DNA — predominantly 150–200 bp fragments — from the acellular fraction of blood. This process differs fundamentally from genomic DNA extraction, which targets cellular material.
The outcome is a purified, concentrated cfDNA sample suitable for highly sensitive downstream assays including next-generation sequencing (NGS), digital droplet PCR (ddPCR), or quantitative PCR (qPCR). The key challenge is working with low-abundance, highly fragmented starting material.
Why Plasma, Not Serum?
Plasma preparation avoids the clotting step required for serum collection. During clotting, leukocytes lyse and release high-molecular-weight genomic DNA that contaminates the sample. Research confirms that EDTA-plasma remains the gold standard for cfDNA analysis, showing minimal DNase activity and low genomic DNA contamination compared to serum.
Genomic DNA contamination matters because it dilutes the tumour-derived or fetal cfDNA signal — directly reducing assay sensitivity in oncology liquid biopsy or NIPT, where variant allele fractions can fall below 0.1%.
The practical implications for sample handling are clear:
- Use EDTA-coated tubes and process plasma within 2 hours of collection to minimise leukocyte lysis
- Perform two-step centrifugation (300–400 × g, then 1,600–3,000 × g) to remove residual cellular debris
- Avoid serum tubes entirely when cfDNA integrity is the priority
Why cfDNA Extraction Matters in Clinical and Research Workflows
The Low Abundance Problem
Healthy individuals carry 1-10 ng/mL of cfDNA in plasma (median ~2.9 ng/mL), while cancer patients show elevated but still modest concentrations:
- Stage I-III cancer: mean 12.6 ng/mL
- Stage I-IV cancer: mean 21.8 ng/mL
- Liver cancer (highest): mean 46.0 ng/mL
Even small extraction losses can push samples below the minimum input requirements for library preparation, which typically require 20 ng of cfDNA for standard NGS workflows.
These yield constraints become even more consequential when you factor in the sensitivity demands of specific clinical applications.
Application-Specific Sensitivity Requirements
Different liquid biopsy applications impose distinct thresholds — and extraction quality determines whether those thresholds are met:
| Application | Key Threshold | Extraction Impact |
|---|---|---|
| Oncology (ctDNA) | VAF <0.01% in early-stage; 0.1–10% in metastatic | Purity sets the floor for low-VAF mutation detection |
| Prenatal testing (NIPT) | Fetal fraction 2–4% minimum; ~10–15% at 10–20 weeks | Poor extraction raises no-call rates, particularly in obese patients (fetal fraction drops ~1% per 10 kg maternal weight gain) |
| Transplant monitoring | Donor-derived cfDNA threshold: 1.0% for kidney rejection | Methods must preserve low-abundance donor DNA without dilution |

What Goes Wrong Without Optimised Extraction
- Genomic DNA contamination inflates background noise and lowers ctDNA fraction detectability
- Low cfDNA yield causes failed library preparation or insufficient template for PCR
- Inconsistent extraction across samples produces non-comparable results in longitudinal monitoring
- Roughly 55% of plasma cfDNA originates from white blood cells — leukocyte lysis during improper handling releases genomic DNA that dilutes the mutant allele fraction below detection thresholds
How cfDNA Extraction from Plasma Works
The complete workflow follows this sequence:
- Blood collection
- Plasma separation
- cfDNA isolation
- Elution
- Quantification and quality assessment
- Downstream analysis
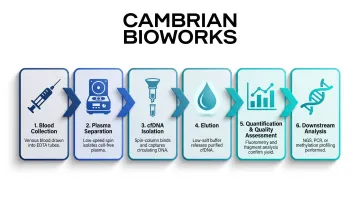
Each upstream step impacts the quality of final cfDNA. Extraction methods rely on selective binding of short DNA fragments (50-700 bp) to a solid-phase substrate — either silica membrane or magnetic beads — followed by washing to remove proteins, lipids, and cellular debris.
Step 1: Blood Collection and Plasma Preparation
Recommended collection protocol:
- EDTA K2 tubes (not serum tubes)
- Process within 2-4 hours for standard EDTA tubes
- Use stabilising tubes (Streck BCT, Roche BCT) if processing will be delayed beyond 4-6 hours
Double centrifugation regimen:
The standard protocol requires two sequential spins:
- First spin: 800-1,600 × g for 10 minutes at 4°C to pellet cells
- Second spin: 14,000-16,000 × g for 10 minutes at 4°C to remove residual cell debris
Skipping the second centrifugation is a leading cause of genomic DNA contamination in downstream assays. EDTA tubes show a 118-fold increase in genomic DNA contamination by day 7 when processing is delayed, compared to only 2.5-fold for stabilising tubes like Roche BCT.
Step 2: cfDNA Isolation — Column vs. Magnetic Bead Methods
Two dominant extraction chemistries exist, each with distinct performance trade-offs:
| Feature | Silica Membrane (Column) | Magnetic Bead |
|---|---|---|
| Binding mechanism | Chaotropic conditions | Bead-surface affinity |
| Total DNA yield | Higher absolute yield | ~3× lower total yield |
| Short fragment enrichment | Lower selectivity | 2.2× more short copies per ng |
| Preferred fragment range | Broad (may co-extract gDNA) | 90-150 bp ctDNA-enriched |
| Automation compatibility | Moderate | High |
| Best application | High-yield research workflows | Oncology / low-allele-fraction assays |
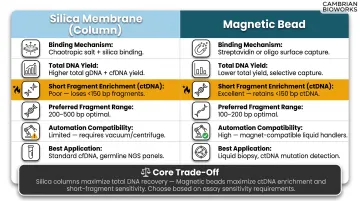
The core trade-off is total yield versus ctDNA enrichment. For liquid biopsy and oncology applications, magnetic bead methods deliver functionally superior input even at lower nanogram quantities — because the fragments recovered are the diagnostically relevant ones.
This selectivity is why automated magnetic bead platforms have gained traction in clinical cfDNA workflows. Cambrian Bioworks' Manta, for example, uses magnetic bead technology with CE-IVD certification to complete cfDNA extraction from plasma in approximately 27 minutes. The system processes 1-32 samples per run using pre-filled cartridges, with flexible input volumes from 100 µL to 4 mL plasma — removing batching pressure without compromising fragment quality.
Step 3: Elution, Quantification, and Quality Assessment
Elution volume decisions: Lower volumes (40-60 µL) concentrate cfDNA but may reduce recovery. Standard recommendations range from 50-60 µL for most clinical applications.
Post-extraction quality control includes:
- Quantification: ddPCR (most sensitive) or fluorometric assays (Qubit) — note that only approximately 49% of extracted cfDNA is amplifiable, meaning Qubit measurements overestimate functional template quantity
- Fragment size profiling: TapeStation or Bioanalyzer to confirm the mononucleosomal peak at approximately 167 bp
- Genomic DNA contamination assessment: Long-to-short fragment ratios using qPCR assays (e.g., 445 bp amplicon vs. 74 bp amplicon) to detect high-molecular-weight DNA carryover
Key Factors That Affect cfDNA Extraction Quality and Yield
Inputs and Pre-Extraction Variables
Sample material type: Always use EDTA plasma, never serum. Serum preparation causes clotting-induced leukocyte lysis and genomic DNA contamination.
Collection tube choice: Standard EDTA tubes require processing within 4-6 hours. Cell-stabilising tubes (Streck, Roche BCT) extend the window to 5-14 days and prevent the dramatic genomic DNA contamination seen with delayed EDTA processing.
Time from blood draw to centrifugation: Processing delays allow cell lysis. EDTA tubes show 118-fold genomic DNA increase by day 7 versus 2.5-fold for Roche BCT.
Freeze-thaw cycles: Up to 3 freeze-thaw cycles do not significantly affect cfDNA stability. Aliquot plasma into single-use volumes to stay within this validated range.
Extraction Method and Technology
Silica column methods: Yield higher absolute cfDNA quantities but co-extract longer fragments, increasing genomic DNA contamination risk.
Magnetic bead methods: Show more consistent inter-run reproducibility and lower variance. Research demonstrates higher short-to-medium fragment ratios and increased variant allele fractions, making them preferable for oncology applications where ctDNA enrichment matters more than total yield.
Method choice should align with downstream application requirements — size selectivity matters critically for fragment length-based analyses.
Input Volume and Elution Parameters
Input plasma volume: The International Society for Liquid Biopsy (ISLB) recommends 4 mL for routine cfDNA applications and 8-20 mL for minimal residual disease detection due to extremely low ctDNA fractions.
Higher input volumes increase yield but may introduce more contaminants. Elution volume and incubation conditions during elution affect final concentration.
Automation and Operator Variability
Manual extraction introduces inter-operator and inter-day variability that confounds longitudinal cfDNA measurements. Automated platforms standardise lysis, wash, and elution steps, reducing technical variability to levels comparable to biological inter-individual differences.
Cambrian Bioworks' Manta platform, for instance, is designed around this principle. Key workflow features include:
- Pre-filled cartridge system that removes manual reagent preparation
- Touchscreen-guided protocol steps that reduce operator-dependent variation
- Flexible run sizes from 1-32 samples, eliminating batching pressure
- Compatibility with oncology ctDNA profiling, prenatal testing, and other cfDNA workflows

Downstream Application Requirements
Validate extraction method performance for each specific clinical application rather than assuming universal suitability.
- NGS-based ctDNA profiling demands higher purity and fragment integrity
- ddPCR-based mutation quantification tolerates more inhibitors
Common Issues, Limitations, and When to Reconsider Your Approach
Higher Yield ≠ Better Results
Labs often assume higher cfDNA yield automatically means better performance. This is misleading. Genomic DNA contamination artificially inflates quantification while reducing mutation allele fraction detectability. Every extraction should include fragment size profiling and contamination checks, not just total yield measurement.
Over-Reliance on a Single Method
Labs sometimes apply one extraction kit across all sample types and clinical applications without validating performance for each use case. A protocol optimised for 1 mL plasma in research may perform inconsistently on 4 mL plasma from cancer patients with abnormal cfDNA concentrations.
When Plasma cfDNA May Not Be Optimal
Some sample types and clinical scenarios fall outside what plasma extraction can reliably address:
- Severely haemolysed samples: Cell lysis before processing introduces genomic DNA contamination that extraction optimisation cannot correct
- Incompatible collection tubes: Blood collected in serum tubes or EDTA-free tubes cannot be salvaged
- Organ-specific cancers with better alternative matrices: Cerebrospinal fluid for gliomas achieves 96.5% ctDNA detection versus 85.4% in plasma; urine cfDNA for bladder cancer reaches 86.7% sensitivity and 99.3% specificity
Signal Below the Noise Floor
For very early-stage cancers or minimal residual disease monitoring, cfDNA abundance may be so low that even high-efficiency extraction cannot yield enough template for reliable variant calling. Consider:
- Increasing input volume to 8–20 mL
- Using carrier-free extraction protocols
- Switching to more sensitive quantification methods (ddPCR over qPCR)
- Consulting published detection thresholds for your specific assay to determine if the clinical question is within reach
Frequently Asked Questions
How do you extract cell-free DNA from plasma?
Separate plasma via double centrifugation (800-1,600 × g then 14,000-16,000 × g) from EDTA blood within 4-6 hours of collection. Use a silica column or magnetic bead-based kit to selectively bind short cfDNA fragments, wash away contaminants, and elute in 40-60 µL for downstream analysis.
How do you quantify and assess the quality of cell-free DNA?
Quantify using ddPCR (most sensitive) or fluorometric methods like Qubit. Assess quality via fragment size profiling on TapeStation or Bioanalyzer to confirm the ~167 bp mononucleosomal peak. Check contamination using long-to-short fragment ratios (e.g., 445 bp vs. 74 bp qPCR assays) to rule out genomic DNA carryover.
How much cell-free DNA is typically found in plasma?
Healthy individuals show 1-10 ng/mL (median ~2.9 ng/mL). Cancer patients have elevated levels: stage I-III mean 12.6 ng/mL, stage I-IV mean 21.8 ng/mL. tumour-derived ctDNA may represent a very small fraction of total cfDNA, so even minor extraction losses can compromise downstream detection.
How accurate is cell-free DNA testing?
Well-validated workflows using optimised extraction achieve high specificity for ctDNA mutation detection. Accuracy declines with poor extraction quality, tumour fraction below 1%, or genomic DNA contamination from delayed processing or haemolysis.
How late in pregnancy can you perform cfDNA testing?
Non-invasive prenatal testing can be performed from 9-10 weeks gestation onwards, with fetal fraction typically reaching 10-15% at 10-20 weeks. Optimal clinical windows start from 10-12 weeks, when fetal fraction reliably exceeds the 2-4% minimum threshold required for accurate results.
Can you extract cell-free DNA from plasma?
Yes, plasma is the gold standard matrix for cfDNA extraction. It is strongly preferred over serum because the clotting step in serum preparation causes leukocyte lysis and genomic DNA contamination. EDTA-plasma shows minimal DNase activity when processed correctly, preserving cfDNA integrity.