Introduction
Cancer outcomes hinge on one critical factor: detection timing. Patients diagnosed at Stage I have significantly better prognoses than those identified at Stage III or IV, when treatment options narrow and survival rates drop sharply. Circulating tumor DNA (ctDNA)-based liquid biopsy is reshaping how clinicians detect, monitor, and respond to cancer at the molecular level — offering a minimally invasive window into tumor biology that complements traditional imaging and tissue analysis.
That clinical promise comes with a real technical challenge: ctDNA is present at extremely low concentrations, often representing less than 1% of total cell-free DNA (cfDNA) in early-stage disease. Method selection, sample quality, and workflow precision are therefore critical for reliable results.
Demand for non-invasive cancer detection has driven rapid market growth — according to a 2023 Grand View Research report, the global liquid biopsy market was valued at $4.27 billion and is projected to grow at a 16.7% CAGR through 2030.
This guide walks through what ctDNA is, its clinical applications, the major detection methods from PCR to next-generation sequencing (NGS), and the pre-analytical factors labs need to manage before any downstream analysis.
TL;DR
- ctDNA is tumor-derived DNA circulating in blood, providing a minimally invasive view of cancer biology
- Clinically applied across early detection, treatment monitoring, and resistance mutation tracking
- ddPCR delivers high sensitivity and rapid results when targeting known mutations
- NGS covers comprehensive multi-gene panels and surfaces novel variants in a single run
- Fragmentomics and methylation analysis broaden detection beyond known mutation targets
What Is ctDNA and How Is It Different from cfDNA?
Circulating tumor DNA (ctDNA) is the tumor-specific fraction of cell-free DNA (cfDNA) found in blood plasma. It is shed by cancer cells through apoptosis, necrosis, or active secretion. While all ctDNA is cfDNA, not all cfDNA is ctDNA, and that distinction directly shapes assay design and interpretation.
The clinical value of ctDNA lies in the tumor-specific alterations it carries. These molecular signatures can be detected from a blood draw alone, enabling real-time monitoring without a tissue biopsy:
- Point mutations — single-nucleotide variants tied to driver oncogenes or resistance mechanisms
- Copy number variations — gene amplifications or deletions reflecting genomic instability
- Methylation patterns — epigenetic changes that can indicate tissue of origin
- Chromosomal rearrangements — structural variants including fusions relevant to targeted therapy selection
ctDNA concentrations vary significantly based on tumor type, stage, and treatment status. In early-stage disease, ctDNA often represents less than 1% of total cfDNA, placing extreme sensitivity demands on detection methods. Advanced-stage cancers typically shed more ctDNA, making detection easier but also indicating disease progression.
Clinical Applications of ctDNA Detection
ctDNA offers a dynamic, real-time picture of tumor biology that imaging and tissue biopsies cannot match, including the ability to capture tumor heterogeneity across multiple disease sites simultaneously.
Four primary clinical use cases:
- Cancer diagnosis and molecular profiling before treatment — identifying actionable mutations to guide therapy selection
- Monitoring treatment response by tracking ctDNA level changes over time
- Detecting minimal residual disease (MRD) post-surgery or post-treatment to identify recurrence risk early
- Identifying acquired resistance mutations to guide therapy switching when first-line treatments fail

These applications are well-established in late-stage disease, but ctDNA has real limitations worth understanding before clinical deployment.
Detection is less reliable in early-stage cancers with low tumor burden. Small or localized tumors below 1 cm, and cancer types with naturally low shedding rates — such as kidney and thyroid cancers — present ongoing challenges. Detection sensitivity varies significantly by stage: approximately 50–75% in Stage I disease compared to >90% in Stage IV across many cancer types.
For this reason, ctDNA testing performs best as a complement to — not a replacement for — imaging and tissue analysis.
ctDNA Detection Methods: A Complete Breakdown
No single ctDNA detection method is universally optimal. Method selection depends on whether the clinical goal is targeted mutation tracking, comprehensive genotyping, epigenetic profiling, or MRD surveillance. Three major categories dominate the field: PCR-based methods, NGS-based methods, and emerging approaches.
PCR-Based Methods (qPCR, ddPCR, and BEAMing)
PCR-based methods — including allele-specific qPCR, digital PCR, droplet digital PCR (ddPCR), and BEAMing — are designed to detect known, predefined mutations with very high sensitivity.
In ddPCR, the sample is partitioned into thousands of droplets, enabling absolute quantification of target molecules without standard curves. This makes it especially effective for detecting rare variants in a wild-type background — some assays achieve detection limits below 0.01% variant allele frequency.
Key advantages:
- High sensitivity for low-abundance mutations
- Faster turnaround times compared to NGS (typically 4-6 hours)
- Simpler workflows with lower bioinformatics demands
- Generally lower per-test cost — more scalable for repeat monitoring in resource-limited or decentralized settings
The core limitation: PCR-based methods require prior knowledge of the target mutation, making them unsuitable for discovery or de novo genotyping.
BEAMing (Beads, Emulsion, Amplification, Magnetics) is a variant of digital PCR that uses magnetic bead-coupled PCR for ultra-high sensitivity mutation detection. Researchers and select clinical programs have used it for monitoring colorectal cancer mutations, though it remains less widely adopted than ddPCR due to higher complexity.
NGS-Based Methods (Targeted Panels and Whole-Genome Approaches)
Next-generation sequencing (NGS)-based ctDNA methods — from targeted gene panels to whole-exome or whole-genome sequencing — allow comprehensive, hypothesis-free genotyping of ctDNA. They identify actionable mutations, copy number alterations, gene fusions, and structural variants across many genes simultaneously.
Tradeoffs:
NGS delivers breadth of detection and suits multi-gene profiling, novel resistance mutation identification, and tumor-informed MRD assays. The tradeoffs versus PCR:
- Requires significant bioinformatics infrastructure
- Longer turnaround times (typically 7-14 days)
- Higher cost per test
- Demands error-correction strategies — unique molecular identifiers (UMIs) or duplex sequencing — to reach the sensitivity needed for low-frequency ctDNA
Tumor-naïve vs. tumor-informed NGS assays:
- Tumor-naïve assays use off-the-shelf gene panels covering common cancer mutations (e.g., 50-500 genes). Preferred when tissue samples are unavailable or for initial molecular profiling
- Tumor-informed assays are personalized, patient-specific panels designed after sequencing the primary tumor. They track dozens to hundreds of patient-specific variants simultaneously, dramatically improving MRD detection sensitivity to levels as low as 1 ctDNA molecule in 100,000 cfDNA molecules
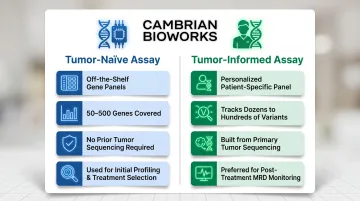
Tumor-informed approaches are preferred clinically for post-treatment MRD monitoring, while tumor-naïve panels are used for treatment selection and resistance monitoring.
Emerging Detection Approaches
Three additional technologies are expanding what ctDNA analysis can detect — moving beyond known mutations into fragmentation patterns, epigenetic signatures, and long-read sequencing.
Fragmentomics analyzes ctDNA fragment length patterns, nucleosome positioning, and end-motif signatures to infer tumor presence and cancer type — without requiring knowledge of specific mutations. Tumor-derived cfDNA exhibits distinct fragmentation patterns compared to normal cfDNA, reflecting differences in chromatin structure and cell death pathways. This approach shows promise for multi-cancer early detection, with studies demonstrating the ability to distinguish cancer types based on fragment profiles alone.
DNA methylation analysis takes an epigenetic approach: tumor-derived ctDNA carries tissue- and cancer-specific methylation signatures that can distinguish cancer type and origin. Methylation-based multi-cancer early detection tests have demonstrated detection rates of 67.6% across 12 cancer types in validation studies, with specificity exceeding 99%.
Third-generation (long-read) sequencing enables direct methylation and structural variant detection from a single molecule, bypassing PCR amplification bias. All three approaches are currently most active in research settings, but regulatory validation efforts — particularly for methylation-based tests — are already underway.
Why Pre-Analytics Matter: The Step That Determines ctDNA Detection Success
The pre-analytical phase — blood collection, tube type, processing time, and cfDNA extraction — is the most overlooked yet most consequential factor in ctDNA testing. Errors here cause cfDNA degradation, genomic DNA contamination, or insufficient yield, which directly reduces the sensitivity and reliability of every downstream detection method.
Pre-analytical variability can account for up to 50-70% of total assay variation in cfDNA testing, undermining even the most advanced detection platforms.
Key pre-analytical considerations:
- Blood collection tube selection: Standard EDTA tubes must be processed within 4–6 hours to prevent white blood cell lysis and genomic DNA contamination. Cell-stabilising tubes (such as Streck or PAXgene) extend stability to 7–14 days at room temperature.
- Time-to-processing: Delay between blood draw and plasma separation degrades ctDNA yield and increases genomic DNA contamination.
- Centrifugation protocol: Double-spin protocols (two sequential centrifugation steps) are recommended to minimise cellular contamination of plasma.
The extraction step is equally decisive. It must maximise cfDNA yield while minimising co-purification of high-molecular-weight genomic DNA. Inefficient extraction directly reduces the detectable ctDNA fraction, producing false negatives even when ctDNA is biologically present.
Automated extraction systems address this by reducing operator-dependent variability and improving reproducibility across samples and operators. Standardising this workflow is essential for serial monitoring (such as MRD tracking over treatment cycles) and for cross-lab consistency in multi-site studies. Without it, an apparent rise in ctDNA levels may reflect a change in centrifuge speed or extraction kit lot — not a change in the patient's disease.
How Cambrian Bioworks Can Help
Reliable ctDNA detection starts before the assay — it starts at extraction. The Manta cfDNA extraction system by Cambrian Bioworks automates this step with a ~55-minute protocol, minimising genomic DNA contamination and maximising cfDNA yield from plasma samples. CE-IVD certified under EU IVDR (2017/746) and ISO 13485 quality-managed, Manta is built for clinical use from the ground up.
Key features:
- Processes 1–32 samples per run with random-access capability
- Compact design (43.5 × 28.2 × 26.2 cm) fits inside a biosafety cabinet or PCR hood
- Pre-filled cartridges eliminate manual reagent pipetting, reducing operator variability
- Remote monitoring capability ensures consistency across runs
- 5–20× higher cfDNA yield compared to leading competitors, using half the plasma volume (1 mL vs. 2 mL)

Pre-analytical consistency is one of the hardest variables to control in ctDNA workflows. Pre-filled cartridges and automated liquid handling remove the manual steps where reproducibility most often breaks down.
Manta is trusted by oncology labs and cancer centres including Tata Memorial Hospital and ACTREC, and is designed for labs that require consistent, audit-ready cfDNA quality as the foundation for any ctDNA detection method, whether the downstream method is PCR-based or NGS-based.
Frequently Asked Questions
Frequently Asked Questions
What counts as early detection for cancer?
Early detection refers to identifying cancer before symptoms appear or before it spreads beyond its original site — typically Stage I or II — where treatment is most effective and survival rates highest. ctDNA-based liquid biopsy can detect molecular signals of cancer weeks to months before imaging finds a visible tumour.
Is cancer curable if detected early?
Many cancers have significantly higher cure rates when caught early. Five-year survival rates for colorectal cancer exceed 90% at Stage I but drop to approximately 14% at Stage IV. Early detection expands treatment options and reduces the need for aggressive intervention.
What is the difference between ctDNA and cfDNA?
cfDNA (cell-free DNA) refers to all DNA fragments circulating in the bloodstream from any cell type, while ctDNA is specifically the subset of cfDNA shed by tumour cells. ctDNA carries cancer-specific genetic alterations, making it the clinically relevant fraction for oncology testing.
Which ctDNA detection method is most sensitive?
ddPCR delivers among the highest sensitivity for detecting known mutations, with limits of detection below 0.01% in optimised assays. For unknown or comprehensive mutation profiling, error-corrected NGS achieves comparable sensitivity but requires greater infrastructure and bioinformatics support.
What are the main limitations of ctDNA testing?
Key limitations include reduced sensitivity in early-stage disease (50–75% at Stage I vs. >90% at Stage IV), lower reliability in cancers with minimal ctDNA shedding (kidney, thyroid), and susceptibility to pre-analytical variability. Results are most reliable when paired with imaging and clinical assessment.
How much blood is needed for a ctDNA liquid biopsy test?
Most ctDNA assays require between 5 and 20 mL of peripheral blood, after plasma separation and cfDNA extraction. The volume required may vary by assay sensitivity, cancer type, and whether a tumour-informed or tumour-naïve approach is being used.


