Introduction
A metastatic lung cancer patient progresses on first-line therapy — but the original tumor site is now unreachable for rebiopsy. This is where cell-free DNA (cfDNA) changes clinical decision-making. Shed into the bloodstream from dying cells, cfDNA can be extracted from a routine blood draw to reveal tumor mutations, treatment resistance markers, and disease burden in real time — without the procedural risks of repeat tissue sampling.
This article covers cfDNA's biological origins, validated clinical applications, and the pre-analytical quality factors — particularly extraction — that determine whether a liquid biopsy delivers actionable results or misleading noise.
TLDR
- cfDNA is fragmented DNA circulating freely in blood plasma, shed by all cell types throughout the body
- ctDNA — the tumor-derived subset of cfDNA — is the primary target biomarker in cancer liquid biopsies
- FDA-approved applications span therapy selection for NSCLC, metastatic breast/prostate cancer, prenatal screening, and transplant monitoring
- Low ctDNA levels in early-stage disease (detection rates as low as 47% in stage I cancers) remain the field's central sensitivity challenge
- cfDNA extraction quality — particularly gDNA contamination control — directly determines assay reliability
What Is Cell-Free DNA? Origins, Biology, and Circulation
cfDNA consists of short, fragmented DNA molecules circulating in the cell-free fraction of biofluids (primarily blood plasma). Unlike DNA inside cells, cfDNA exists as extracellular fragments released by all cell types through normal turnover, apoptosis, necrosis, and active secretion.
Fragment Size and Nucleosome Protection
cfDNA fragments peak at approximately 166 base pairs, reflecting DNA wrapped around histone proteins. This size corresponds to mono-nucleosomal DNA (147 bp around the histone core plus ~20 bp linker DNA). Studies show a characteristic 10 bp periodicity that reflects helical turns around histones, with mono-nucleosomes accounting for 67-80% of total cfDNA in healthy individuals.
ctDNA fragments are characteristically shorter than non-cancerous cfDNA. Cancer patients show enrichment of fragments between 40-150 bp with a shoulder at 145-160 bp. This size difference is analytically exploited: laboratories can enrich for tumor signal by selecting shorter fragments during extraction and library preparation.
Rapid Clearance Enables Real-Time Monitoring
cfDNA clears from circulation quickly, with a half-life of 16 minutes to 2.5 hours depending on the study. Circulating tumor DNA specifically clears in under 1.5 hours. The liver is the primary clearance organ, with Kupffer cells and sinusoidal endothelial cells internalizing and degrading cfDNA. This rapid turnover has two critical implications:
- Serial blood draws can track treatment response within hours to days, unlike protein biomarkers with longer half-lives
- Low-burden tumors release minimal ctDNA into a system that clears it aggressively, creating sensitivity challenges
Cellular Origins of cfDNA
| Source | Mechanism | Fragment Characteristics |
|---|---|---|
| Apoptosis | Caspase-activated cleavage at inter-nucleosomal sites | ~180 bp laddering pattern |
| Necrosis | Non-specific release from dying cells | Large fragments (>10,000 bp) |
| Active secretion | Extracellular vesicles (exosomes, microvesicles) | 100 bp to >2 million bp |
| NETosis | Neutrophil extracellular trap release | Long DNA filaments |
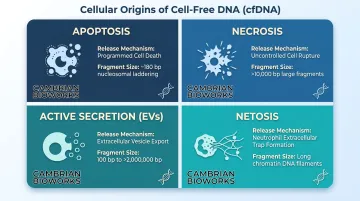

Notably, cancer cells actively secrete heterochromatin segments of 2,000-3,000 bp, and DNA protected within extracellular vesicles is shielded from circulating nucleases, extending its persistence in circulation.
cfDNA vs. ctDNA: Understanding the Key Distinction
ctDNA is the tumor-derived subset of total cfDNA. Healthy individuals carry background cfDNA from normal cell turnover; cancer patients carry additional ctDNA from tumor apoptosis and necrosis. The analytical challenge is detecting the ctDNA signal against this background noise — and the key variable governing that challenge is tumor fraction.
How Tumor Fraction Shapes Detection
Tumor fraction, the proportion of total cfDNA originating from cancer cells, varies from less than 0.01% to over 10% depending on cancer type, stage, and disease burden. A tumor must contain roughly 50 million malignant cells to release detectable DNA into circulation.
Bettegowda et al. (2014) demonstrated stage-dependent detection rates across solid tumors:
- Stage I: 47% detection rate
- Stage II: 55%
- Stage III: 69%
- Stage IV: 82%
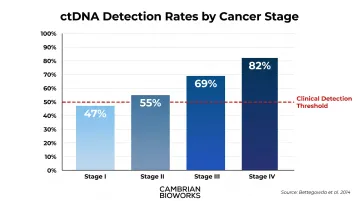
These figures show that nearly half of early-stage cancers have undetectable ctDNA using current technologies. This sensitivity gap is the fundamental reason liquid biopsy cannot yet replace tissue diagnosis in early-stage settings.
Detection Rates Vary Significantly by Cancer Type
Tumor biology adds another layer of variability. ctDNA is detectable in over 75% of advanced cases for several cancer types, including:
- Pancreatic and colorectal cancers
- Ovarian and bladder cancers
- Melanoma and head/neck cancers
Gliomas, by contrast, show detection rates below 10%. The blood-brain barrier limits DNA release into peripheral circulation, making CNS tumors among the most challenging targets for liquid biopsy.
How cfDNA Liquid Biopsy Testing Works
The analytical workflow begins with blood collection and extends through bioinformatic interpretation. Each step introduces potential variability that can degrade test performance.
Pre-Analytical Variables
Blood collection tube selection critically impacts results:
| Tube Type | Processing Window | gDNA Contamination Risk |
|---|---|---|
| EDTA | 4-6 hours | High after 6 hours |
| Streck Cell-Free DNA BCT | Up to 14 days at room temperature | Minimal even after 96 hours |
EDTA requires rapid processing; Streck tubes stabilise cells and prevent genomic DNA contamination from lysing white blood cells, making them ideal for decentralised collection and shipped specimens.
Extraction: The Foundation of Quality
Following plasma separation by double centrifugation, cfDNA extraction determines everything downstream. Inadequate extraction co-isolates high-molecular-weight genomic DNA, diluting ctDNA signal and introducing false positives. High-yield, selective extraction is therefore a foundational requirement — not an optional upgrade.
Purpose-built platforms address this directly. Cambrian Bioworks' Manta (CE-IVD certified, magnetic bead-based) completes cfDNA extraction in approximately 55 minutes, supports 1-4 mL plasma input volumes, and maintains fragment enrichment in the 150-200 bp range critical for tumour signal detection. According to Cambrian Bioworks' published performance data, the system delivers 5-20× higher cfDNA yield compared to leading competitors from just 1 mL of plasma, with minimal gDNA contamination throughout.

Detection Technologies
Major platforms include:
- NGS panels: Detect somatic mutations and copy number variants; Guardant360 CDx covers 55 genes, FoundationOne Liquid CDx covers over 300 genes
- Methylation assays: Classify tissue-of-origin using methylation signatures (for example, the Galleri MCED test)
- Fragmentomics: Analyse fragment size, end motifs, and nucleosome positioning to infer cancer presence and origin
Most commercial panels report results in 7-10 business days, though same-day extraction at the lab level combined with rapid sequencing workflows can accelerate clinical decision-making.
Clinical Applications of cfDNA in Oncology
Companion Diagnostics and Therapy Selection
The most clinically validated use of ctDNA is as a companion diagnostic when tissue is unavailable or biopsy is unsafe. FDA-approved liquid biopsy tests include:
NSCLC indications:
- EGFR exon 19 deletions and L858R mutations for osimertinib, gefitinib, erlotinib
- KRAS G12C for sotorasib and adagrasib
- ALK rearrangements for alectinib
- ROS1 fusions for entrectinib
Metastatic breast cancer:
- PIK3CA mutations for alpelisib (Piqray)
- ESR1 mutations for elacestrant (Orserdu)
Metastatic prostate adenocarcinoma:
- BRCA1/2 and other HRR mutations for PARP inhibitors (olaparib, rucaparib)
These indications share a common feature: high tumor burden in metastatic settings typically produces tumor fractions above 1%, making ctDNA reliably detectable.
Treatment Monitoring and Resistance Detection
Serial ctDNA measurements during therapy enable real-time tracking of tumor dynamics. Two signals drive clinical decisions:
- Rising ctDNA indicates emerging resistance or progression — often before imaging changes appear
- Declining ctDNA confirms therapeutic response and guides continuation of treatment
This pharmacodynamic approach supports adaptive treatment strategies in metastatic disease, allowing oncologists to act on molecular evidence rather than waiting for radiographic confirmation.
Minimal Residual Disease (MRD) Detection
Where on-treatment monitoring tracks active disease, MRD testing looks for residual disease after treatment — detectable by ctDNA but invisible to imaging. The DYNAMIC trial in stage II colon cancer demonstrated proof-of-concept: ctDNA-guided management halved chemotherapy use (15% vs. 28%) while achieving non-inferior 2-year recurrence-free survival (93.5% vs. 92.4%).
That said, NCCN and ASCO do not recommend routine ctDNA-based MRD testing outside clinical trials for most solid tumors. The evidence base remains investigational for most cancers. Payer coverage is similarly limited: Medicare covers select tests like Guardant Reveal for CRC surveillance as of 2025, but broader reimbursement awaits prospective interventional trials demonstrating outcome improvement.
Multi-Cancer Early Detection (MCED)
MCED tests use methylation signatures to detect multiple cancer types from a single blood draw. The PATHFINDER 2 study (n=25,578) demonstrated Galleri's performance:
- Specificity: 99.6%
- Positive predictive value: 61.6%
- Cancer signal origin accuracy: 92%
- Episode sensitivity for 12 deadliest cancers: 73.7%
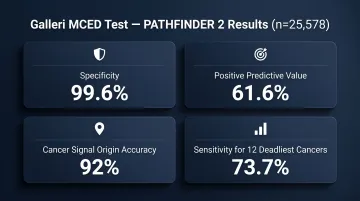
While these metrics support feasibility, MCED tests lack insurance coverage and require prospective interventional studies demonstrating mortality benefit before widespread screening use is justified. A positive MCED result triggers diagnostic workup—it does not establish a cancer diagnosis.
cfDNA Beyond Oncology
Prenatal Testing (NIPT)
Non-invasive prenatal testing is cfDNA's most clinically mature application outside oncology. Fetal cfDNA circulates in maternal plasma from early gestation, enabling detection of chromosomal aneuploidies. Meta-analyses show detection rates exceeding 99% for trisomy 21, 98% for trisomy 18, and 99% for trisomy 13, with a combined false positive rate of 0.13%.
ACOG recommends cfDNA as "the most sensitive and specific screening test for common fetal aneuploidies in any patient population." Positive results require confirmatory invasive testing (amniocentesis or CVS) before clinical decisions are made.
Organ Transplant Monitoring
Donor-derived cell-free DNA (dd-cfDNA) serves as a non-invasive biomarker for graft injury — when donor cells die, they release DNA distinguishable from recipient cfDNA. Elevated dd-cfDNA levels are associated with acute rejection episodes in kidney, heart, and lung transplants. The AlloSure assay uses a 1.0% threshold for active rejection detection, achieving sensitivity of 59% and specificity of 85%, with a negative predictive value of 84%.
Emerging Applications
Investigational uses include:
- Pathogen-derived cfDNA detection for sepsis and opportunistic infection diagnosis (Karius Spectrum test reports 93.7% sensitivity)
- cfDNA levels and methylation patterns as activity markers in autoimmune disease monitoring
- Tissue damage assessment via cfDNA release following trauma or acute injury
These remain research tools rather than validated clinical diagnostics.
Current Limitations and Analytical Challenges
Sensitivity and the Tumor Fraction Problem
The stage I detection rate of 47% fundamentally limits cfDNA use in early-stage disease. When ctDNA represents a fraction of a percent of total cfDNA, even highly sensitive assays struggle.
Clonal hematopoiesis of indeterminate potential (CHIP) compounds this challenge. Somatic mutations in blood cell precursors—particularly in DNMT3A, TET2, TP53, and ASXL1—create false-positive ctDNA signals. CHIP prevalence is age-related, reaching 10-15% in patients aged 70-80. Without filtering, 76.4% of cfDNA mutations may be CHIP-derived rather than tumor-derived.
Matched leukocyte sequencing identifies mutations present in both plasma and white blood cells, flagging them as CHIP. This approach reduces false positive rates by over 10-fold and is incorporated into leading platforms like Guardant360 CDx and FoundationOne Liquid CDx.
Pre-Analytical Variability
Blood collection tube type, time to plasma separation, freeze-thaw cycles, and extraction method all affect cfDNA yield and integrity. Lack of standardization across labs is a major barrier to multicenter reproducibility and clinical adoption.
The BLOODPAC consortium has published minimum technical data elements to harmonize practices, and clinical practice guidelines specify protocols for tube selection, centrifugation, and storage. CE-IVD certified, validated extraction workflows matter because they eliminate this source of variability.
Clinical Validity vs. Utility Gap
Analytical validity (whether a test measures what it claims) is distinct from clinical utility (whether the result actually improves patient outcomes). Many cfDNA assays have cleared the analytical bar but haven't been proven in prospective interventional trials to change management in ways that improve survival or reduce harm.
This evidence gap has real consequences:
- Payers classify MRD testing as investigational for most solid tumors
- MCED tests lack coverage despite demonstrated feasibility
The Future of cfDNA Liquid Biopsy
Modulating cfDNA Biology to Improve Sensitivity
Rather than solely improving sequencing technology, emerging research explores upstream interventions to increase informative cfDNA molecules before analysis. Martin-Alonso et al. (Science, 2024) demonstrated two approaches:
Liposomal priming agents saturate the mononuclear phagocyte system (Kupffer cells), transiently reducing hepatic clearance. This increased cfDNA by 10-fold at 100 mg/kg and up to 78-fold at 300 mg/kg in healthy mice. In cancer models, priming agents improved small tumor detection from less than 10% to over 75%.
Anti-DNA monoclonal antibodies bind cfDNA to protect from nuclease degradation, improving mutant molecule recovery by a median 19-fold.
Sonobiopsy uses focused ultrasound with microbubbles to locally disrupt the blood-brain barrier or tumor vasculature, enhancing ctDNA release. This approach is particularly relevant for CNS tumors where blood-based detection rates are otherwise below 10%.
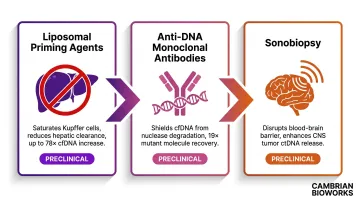
All three approaches remain preclinical. Clinical translation will require toxicity profiling, dose optimisation, and regulatory pathways — but each addresses the sensitivity ceiling that sequencing improvements alone cannot resolve.
AI, Multi-Analyte Integration, and Standardisation
Next-generation liquid biopsy will likely move beyond single-analyte testing, combining cfDNA with other circulating markers in AI-interpreted multi-omic panels. Key analytes under active investigation include:
- Circulating tumor cells (CTCs)
- Exosomes and extracellular vesicles
- Cell-free RNA and microRNA
- Circulating proteins and metabolites
Standardised minimum data elements and reporting requirements will be essential for cross-study comparability and regulatory acceptance as this multi-analyte approach matures.
Frequently Asked Questions
How accurate is a cell-free DNA test?
Accuracy depends heavily on cancer type, stage, and tumor burden. In metastatic settings with high ctDNA shedding, detection rates exceed 80%, but drop to 47% in stage I disease. Specificity exceeds 99% when matched white blood cell sequencing filters CHIP false positives.
What is a cell free DNA test for cancer?
A cfDNA cancer test (liquid biopsy) analyses circulating tumor DNA from a blood sample to identify cancer-associated mutations or methylation patterns. It is used for companion diagnostic testing, treatment selection, and disease monitoring, complementing rather than replacing tissue biopsy.
What happens after a positive cfDNA test?
In a companion diagnostic context, a positive result may directly guide therapy selection. In a screening or MCED context, it requires confirmatory diagnostic workup including imaging and tissue biopsy. A positive cfDNA result alone is not sufficient for a cancer diagnosis.
What is the difference between cfDNA and ctDNA?
cfDNA is all cell-free DNA circulating in blood plasma from any cell type, while ctDNA is specifically the tumor-derived fraction. ctDNA typically makes up a small and variable percentage of total cfDNA and is the clinically informative signal in cancer liquid biopsies.
How much does a cell free DNA test cost?
Costs vary widely by test type (USD-denominated US market benchmarks):
- Prenatal cfDNA (NIPT): $800–$2,000 out-of-pocket
- Guardant360 CDx: $5,000 under US Medicare reimbursement
- Galleri MCED: $949 list price
Coverage in any market depends on whether the test meets payer medical necessity criteria for the indication.
Can cfDNA testing replace tissue biopsy?
No. cfDNA liquid biopsy complements but does not replace tissue biopsy. Tissue remains required for histological diagnosis, and ctDNA testing is recommended only when tissue is unavailable, insufficient, or biopsy is unsafe. A negative cfDNA result does not rule out cancer.