Introduction
Cancer diagnosis has long depended on invasive tissue biopsies—procedures that carry risk, patient discomfort, and logistical complexity. Today, a simple blood draw can reveal tumor DNA, mutations, and treatment resistance in real time—shifting oncology toward faster, less invasive decision-making at every stage of care.
Liquid biopsy combined with circulating tumor DNA (ctDNA) analysis and next-generation sequencing (NGS) is now central to early cancer detection, treatment monitoring, and resistance profiling. But the clinical reliability of these tests depends entirely on pre-analytical quality—particularly cfDNA extraction.
This guide covers the complete liquid biopsy NGS workflow: from blood collection and cfDNA isolation to library preparation, sequencing, and clinical interpretation—with a focus on the variables that determine whether results are clinically trustworthy.
TL;DR
- Liquid biopsy analyzes ctDNA and other biomarkers from blood instead of tissue
- ctDNA represents less than 1% of total cfDNA, requiring ultra-sensitive extraction and sequencing
- NGS enables comprehensive mutation profiling far beyond PCR capabilities
- Poor extraction quality is the leading cause of unreliable NGS results, making pre-analytical steps critical
- Clinical applications include therapy selection, MRD monitoring, and early recurrence detection
What Is Liquid Biopsy, ctDNA, and NGS?
Liquid biopsy is a minimally invasive diagnostic method that analyzes tumor-derived material—including circulating tumor DNA (ctDNA), circulating tumor cells (CTCs), exosomes, and non-coding RNAs—from blood, urine, or cerebrospinal fluid, without requiring tissue.
Understanding cfDNA vs. ctDNA:
Cell-free DNA (cfDNA) is all fragmented DNA circulating in the bloodstream, originating from normal cell turnover. ctDNA is the tumor-specific subset carrying cancer-associated mutations, methylation patterns, and copy number changes—typically representing less than 1% of total cfDNA.
In early-stage cancer, ctDNA can constitute less than 0.1% of cfDNA, while in advanced disease it may exceed 10%. This wide variation means extraction protocols must maximise ctDNA yield while minimising contamination from high-molecular-weight genomic DNA released by white blood cell lysis.
The Role of NGS:
Next-Generation Sequencing (NGS) is the high-throughput sequencing technology used to detect and characterise ctDNA mutations, copy number variants, and methylation profiles from liquid biopsy samples.
Unlike traditional PCR-based methods, NGS delivers the sequencing depth and sensitivity needed to detect variants at allele frequencies below 0.1%—critical for early detection and minimal residual disease monitoring.
How They Connect
Liquid biopsy → cfDNA extraction → ctDNA enrichment → NGS creates a complete non-invasive diagnostic pipeline. Each step is interdependent:
- Poor extraction yields poor sequencing data
- Poor sequencing data leads to missed variants or false results
- gDNA contamination from white blood cell lysis can increase 118-fold within 7 days in standard EDTA tubes, masking the ctDNA signal entirely
This pipeline integrity is why labs invest in validated, high-quality extraction workflows.
Why Liquid Biopsy + ctDNA + NGS Matters for Cancer Diagnostics
Core Advantages Over Tissue Biopsy
Liquid biopsy captures tumor heterogeneity across all sites (primary tumor plus metastases) and can be repeated serially at low risk to the patient. It reflects real-time tumor dynamics, making it well-suited to monitoring treatment response and detecting resistance before clinical progression appears on imaging.
Key advantages over tissue biopsy include:
- Samples all tumor sites simultaneously, capturing spatial heterogeneity missed by single-site biopsy
- Enables serial monitoring without invasive procedures
- Fills the gap when tissue is unavailable — ~20% of metastatic NSCLC patients have insufficient tissue for NGS
- In tissue-insufficient cases, plasma ctDNA-NGS identified actionable alterations in 51% of patients, with a 70% objective response rate on matched targeted therapy
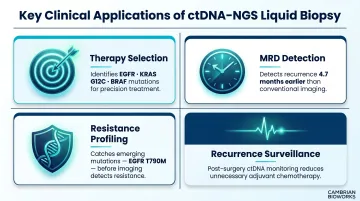
Key Clinical Applications
- Therapy selection: Identifies actionable mutations (EGFR, KRAS G12C, BRAF) to guide precision treatment matching
- MRD detection: Detects recurrence 4.7 months earlier than imaging in resected EGFR-mutant NSCLC (ADAURA trial)
- Resistance profiling: Serial monitoring catches emerging mutations like EGFR T790M before radiographic progression
- Recurrence surveillance: MRD monitoring post-surgery has transformed follow-up care, though sensitivity in Stage I-II disease remains below 30% for many cancer types

Clinical Validation
ctDNA-NGS achieves 70-80% sensitivity versus tissue-based genotyping for advanced NSCLC, with specificity exceeding 98% for EGFR mutations. FDA-approved assays include:
- Guardant360 CDx (74 genes, approved August 2020)
- FoundationOne Liquid CDx (324 genes, expanded May 2023)
- cobas EGFR Mutation Test v2 (real-time PCR, approved June 2016)
All three carry companion diagnostic approvals, meaning they're validated not just for detection but for directly informing treatment decisions.
How ctDNA Diagnosis Through NGS Works—Step by Step
ctDNA diagnosis via NGS involves distinct pre-analytical, analytical, and post-analytical phases. Failures at any one stage cascade into unreliable downstream results. The extraction step is the most common failure point for labs transitioning to liquid biopsy workflows.
Step 1 – Sample Collection and Stabilisation
Blood collection protocols for liquid biopsy require cfDNA-preserving collection tubes—such as EDTA-based or cell-stabilising tubes—to prevent genomic DNA contamination from white blood cell lysis.
Critical timing: EDTA tubes show a 118-fold increase in gDNA contamination by day 7, while stabilising tubes (Streck BCT, Roche BCT) maintain cfDNA integrity for 7-14 days at room temperature.
Samples should be processed within defined windows to preserve ctDNA integrity. Storing blood at 4°C for up to 48 hours preserves cfDNA better than room temperature storage.
Metrics impacted: Sample quality, ctDNA yield, downstream sensitivity.
Step 2 – cfDNA Extraction and Isolation
Plasma separation involves centrifugation steps to remove cells and cellular debris, followed by cfDNA extraction using magnetic bead-based or column-based methods.
Fragment size matters: Normal cfDNA peaks at approximately 167 bp, while tumor-derived ctDNA fragments are consistently shorter at 134-145 bp. Extraction protocols must recover these short fragments while minimising high-molecular-weight genomic DNA contamination.
Magnetic bead-based methods achieve 76-87% cfDNA recovery rates with strong linearity between plasma input volume and recovered DNA.
Automated extraction systems with validated protocols help standardize this critical step and deliver the expected mononucleosomal (~167 bp) and dinucleosomal (~340 bp) fragment distribution.
Cambrian Bioworks' Manta platform addresses the most failure-prone step in the liquid biopsy workflow with:
- CE-IVD certification for clinical-grade cfDNA extraction
- Magnetic bead-based cfDNA extraction in ~55 minutes total workflow time
- Minimal gDNA contamination
- Processing flexibility (1-32 samples per run, no batching pressure)
- Open system compatibility with downstream library prep kits
Typical cfDNA yield from 3-4 mL plasma is approximately 20-30 ng.
Metrics impacted: cfDNA yield, gDNA contamination, fragment size integrity.
Step 3 – Library Preparation
cfDNA is converted into sequencing-ready libraries through end repair, adapter ligation, and optional targeted enrichment (panel-based capture or amplicon-based approaches).
Unique Molecular Identifiers (UMIs): UMIs are short random nucleotide sequences (4-12 bp) ligated to each original DNA molecule before PCR amplification. After sequencing, reads sharing the same UMI are collapsed into a consensus sequence, enabling error correction and detection of very low-frequency variants.
The Guardant360 CDx platform employs UMI-based digital sequencing, achieving analytical calling thresholds of SNV MAF ≥0.001% and indel MAF ≥0.01%. Without UMI-based error correction, the background error rate of standard NGS (approximately 0.1-1%) would preclude detection of variants at these frequencies.
Metrics impacted: Library complexity, coverage uniformity, limit of detection.
Step 4 – NGS Sequencing
Targeted gene panels (most common clinically):
- High depth (10,000-100,000× coverage)
- Lower cost ($800–$4,000 per test)
- Specific mutations in 50-500 genes
- Enables detection of variants below 0.1% MAF
Whole exome sequencing:
- Broader variant landscape (~20,000 genes)
- Shallower depth (200-500×)
- Low sensitivity for variants below 1% MAF
Whole genome sequencing:
- Copy number, structural variants, methylation
- Standard depth (30-60×) very low for rare variants
- Fragmentation patterns (DELFI) offer complementary cancer detection strategy
Ultra-deep sequencing is needed to detect ctDNA variants present at allele frequencies below 1%. The Cristiano et al. DELFI study demonstrated that genome-wide fragmentation patterns could distinguish cancer patients from healthy individuals with 94% AUC, detecting 79% of Stage I-III patients at 98% specificity.
Metrics impacted: Variant detection sensitivity, allele frequency resolution, assay cost.
Step 5 – Bioinformatics and Variant Calling
The bioinformatics pipeline includes:
- Alignment to reference genome
- Duplicate removal (UMI-based deduplication)
- Variant calling with error-suppression algorithms
- Filtering sequencing artefacts vs. true somatic mutations
- Annotation against clinical databases (ClinVar, COSMIC)

Clonal Hematopoiesis (CHIP) Challenge:
CHIP is a major source of false positives. A study analysing 4,324 oncology cfDNA samples found that 30.3% showed evidence of clonal hematopoiesis. The most commonly affected genes—DNMT3A, TET2, PPM1D, and TP53—overlap with genes frequently mutated in solid tumours.
The established approach is paired sequencing of white blood cells (WBCs) alongside cfDNA: variants present in both are classified as CHIP-derived rather than tumour-derived. Machine learning approaches are emerging that can classify CHIP status from cfDNA sequencing alone.
Metrics impacted: False positive rate, clinical interpretation accuracy, reporting confidence.
Step 6 – Clinical Interpretation and Action
Clean bioinformatics output feeds directly into treatment decisions. Detected ctDNA variants are classified (pathogenic, likely pathogenic, variant of uncertain significance) and translated into three core clinical actions:
- Therapy selection: Matching mutations to approved targeted therapies or clinical trials
- Treatment monitoring: Serial ctDNA quantification tracks response or MRD—in advanced NSCLC, plasma ctDNA response preceded radiographic response by a median of 24 weeks
- Recurrence detection: Rising ctDNA levels flag early relapse; in the DYNAMIC trial, ctDNA-guided adjuvant therapy in Stage II colon cancer reduced chemotherapy use from 28% to 15% without compromising recurrence-free survival (93.5% vs. 92.4%)
Metrics impacted: Time to treatment decision, actionability rate, patient outcome relevance.
A Liquid Biopsy ctDNA-NGS Workflow Walkthrough
To make the workflow concrete, this walkthrough follows a patient with advanced NSCLC — the clinical scenario where ctDNA testing has the strongest validation and broadest clinical adoption.
Scenario: A patient with advanced NSCLC has insufficient tissue from a small biopsy for genomic profiling. The oncologist orders a liquid biopsy.
Step-by-step:
- Blood collection: Blood is collected into cfDNA-stabilising tubes within 2 hours of draw. Common pitfall: delays in processing degrade ctDNA.
- Plasma separation: Two-step centrifugation removes cells and debris. Common pitfall: skipping QC steps misses gDNA contamination.
- cfDNA extraction: Automated magnetic bead-based system extracts cfDNA in ~55 minutes with minimal gDNA contamination.
- Library preparation: UMI-tagged libraries are prepared for targeted NGS panel analysis.
- Sequencing: Ultra-deep sequencing (10,000× coverage) detects an EGFR exon 19 deletion at 0.4% allele frequency—below the limit of detection for standard PCR. Common pitfall: shallow sequencing misses low-frequency EGFR T790M resistance mutations.
- Interpretation: The mutation informs the decision to start osimertinib therapy.
- Monitoring: Serial monitoring over 3 months shows ctDNA clearance, confirming treatment response before imaging shows tumour shrinkage.
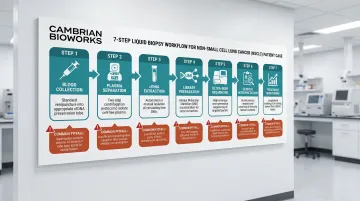
Key takeaway: The reliability of this outcome depends entirely on pre-analytical quality and sequencing depth. A lapse at any single step — delayed plasma separation, inadequate sequencing coverage, or insufficient QC — can convert a clinically actionable result into a missed or false finding.
How Cambrian Bioworks Supports Your Liquid Biopsy NGS Workflow
Cambrian Bioworks provides precision extraction solutions designed for the pre-analytical challenges of liquid biopsy and ctDNA diagnostics.
Manta Platform Key Features:
- CE-IVD certified under IVDR 2017/746 for clinical-grade cfDNA extraction
- ISO 13485 quality management for medical device manufacturing
- Completes cfDNA extraction in ~55 minutes total (~30 minutes extraction)
- Preferentially recovers short ctDNA fragments (150-200 bp) with minimal gDNA contamination
- Processes 1-32 samples per run — no batching pressure
- Open system compatibility for integration with downstream library prep kits
- Remote monitoring for workflow oversight
These technical specifications translate directly into measurable performance gains at the bench.
Performance Advantages:
- Maximises cfDNA yield from plasma (100-1,365 ng per mL in validation studies)
- Preserves short fragment integrity critical for oncology workflows
- Pre-filled cartridge system minimises hands-on time and contamination risk
- Automated magnetic bead-based technology ensures run-to-run reproducibility

That combination of yield, integrity, and reproducibility underpins clinical confidence — which is why Manta has earned adoption across regulated diagnostic settings.
Clinical-Grade Reliability:
Trusted by 67+ labs globally across oncology, molecular diagnostics, and research settings in 5+ countries, with customers including Tata Memorial Hospital, ACTREC, and Strand Life Sciences.
Manta targets the pre-analytical failure points that most often compromise NGS sensitivity: gDNA contamination, low ctDNA input, and inconsistent extraction yields. That makes it a practical fit for both high-throughput clinical labs and smaller research settings building or scaling liquid biopsy programmes.
Frequently Asked Questions
What is a NGS liquid biopsy?
A NGS liquid biopsy is a minimally invasive test that uses next-generation sequencing to analyse circulating tumour DNA (ctDNA) or other biomarkers isolated from blood or bodily fluids, enabling detection of cancer-associated mutations, copy number changes, and other genomic alterations without a tissue biopsy.
Is NGS suitable for liquid biopsy?
Yes, NGS is highly suitable and increasingly the preferred method for liquid biopsy analysis. It offers comprehensive, high-throughput mutation profiling at the sequencing depths needed to detect low-frequency ctDNA variants — something PCR-based methods cannot replicate for broad panel testing.
What are the FDA approved liquid biopsy assays?
Several liquid biopsy assays have received FDA approval, including the Guardant360 CDx (for EGFR mutations in NSCLC), FoundationOne Liquid CDx (comprehensive pan-cancer panel), and the cobas EGFR Mutation Test v2. Check the FDA website for current approvals, as new assays are regularly approved.
Does insurance pay for a liquid biopsy?
Coverage varies by payer, indication, and country. In the US, Medicare and many private insurers cover FDA-approved companion diagnostic tests for specific cancers like NSCLC — though 70% of private payers restrict coverage to cases where tissue biopsy is unavailable or contraindicated.
What is ctDNA and how is it different from cfDNA?
cfDNA (cell-free DNA) refers to all fragmented DNA circulating in the bloodstream, originating from normal cell turnover. ctDNA (circulating tumour DNA) is the tumour-derived subset of cfDNA that carries cancer-specific mutations and typically constitutes less than 1% of total cfDNA—making sensitive extraction and sequencing methods essential for reliable detection.
What are the main limitations of liquid biopsy NGS?
Key limitations include:
- Low ctDNA abundance in early-stage disease increases false negative rates
- Clonal haematopoiesis introduces false positives unrelated to tumour mutations
- Lack of standardised protocols across labs affects reproducibility
- High per-test costs ($1,000–$5,000) and complex bioinformatics limit routine adoption
Detection sensitivity drops below 30% for Stage I-II tumours in multi-cancer screening contexts.


