FFPE samples are the most widely used clinical specimens in oncology, but extracting sequencing-grade DNA from them is often unreliable due to variability in fixation and processing.
Our kit uses bead-based chemistry to consistently recover high quality DNA even at low tumor content (~10%), with broad fragment coverage (200 bp–48.5 kbp) and strong DIN values, making it ready for NGS and PCR workflows.
Consistency across
FFPE variability
Optimized to handle variability in paraffin and formalin quality across regions
The kit delivers reliable DNA extraction across variable FFPE blocks
Performance preview
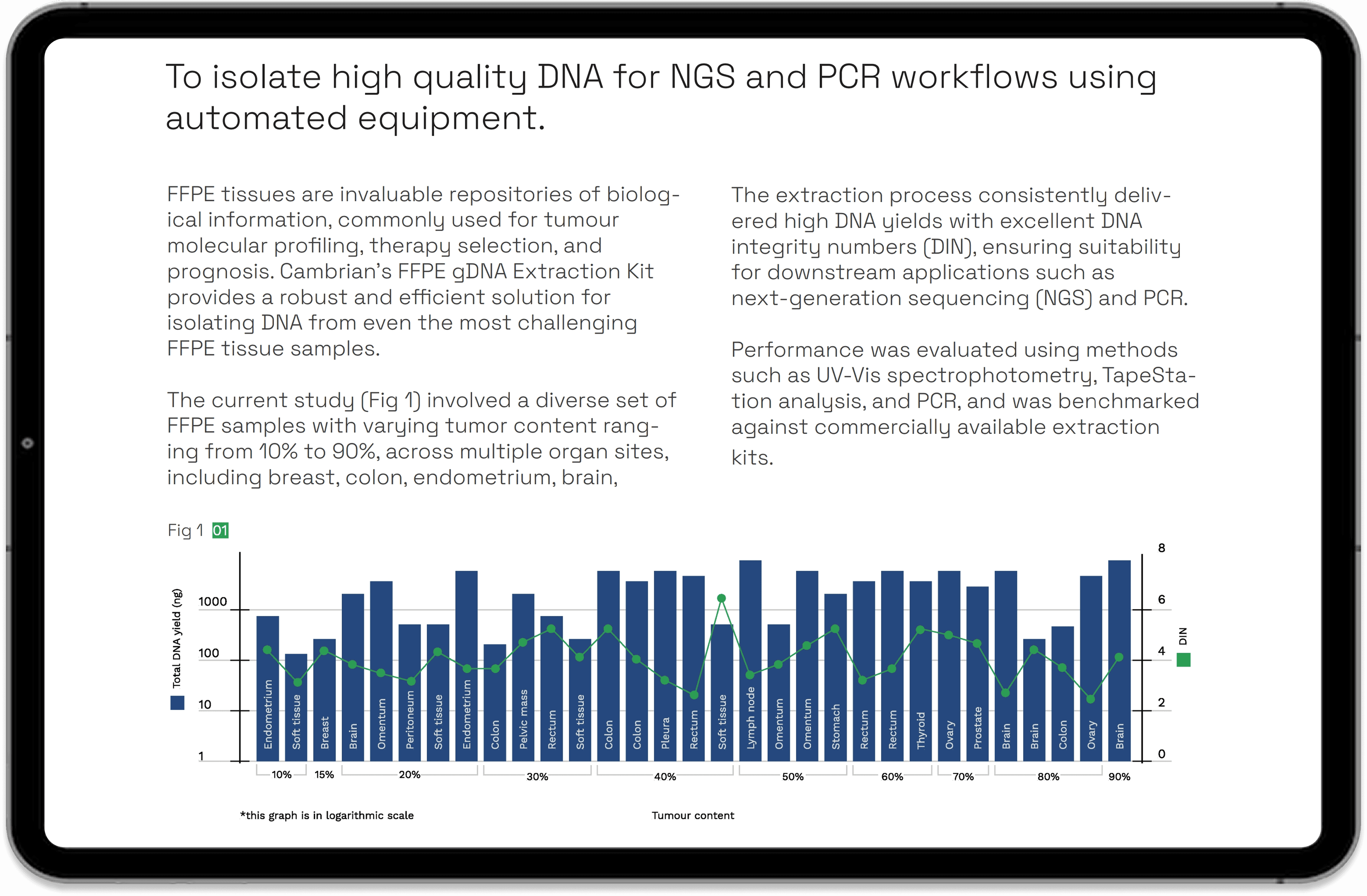
See how the Cambrian FFPE DNA Extraction Kit performs on real-world tumour samples. This application note highlights DNA yield, integrity (DIN), and sequencing readiness across diverse FFPE tissues demonstrating reliable performance for NGS and PCR workflows.
Ordering information
Make FFPE extraction reliable
Handle variability with confidence and get consistent DNA from every block.




