Introduction
Completing cancer treatment and achieving apparent remission doesn't guarantee disease eradication. Residual cancer cells — too scarce for traditional pathology or imaging to detect — can remain hidden in the body and ultimately trigger relapse. This clinical reality has driven the development of measurable residual disease (MRD) detection, a set of highly sensitive molecular assays capable of identifying malignant cells at concentrations far below the threshold of conventional microscopy.
MRD monitoring has reshaped how oncologists manage blood cancers such as acute myeloid leukemia (AML), B-cell acute lymphoblastic leukemia (ALL), chronic myeloid leukemia (CML), and multiple myeloma. Instead of waiting for clinical symptoms or radiographic evidence of recurrence, clinicians can now make treatment decisions based on molecular-level data that reveals the true depth of remission.
Beyond blood cancers, MRD is also gaining traction in solid tumors through circulating tumor DNA (ctDNA) liquid biopsy — extending the reach of molecular surveillance to a much broader patient population.
This guide covers the principal MRD detection methods in clinical use today — flow cytometry, PCR-based assays, next-generation sequencing (NGS), and ctDNA liquid biopsy — examining how they differ in sensitivity, application, sample requirements, and turnaround time.
It also addresses what labs need to consider when selecting or optimizing MRD testing workflows, from infrastructure and expertise to the critical role of high-quality sample preparation.
TL;DR
- MRD is residual cancer cells surviving treatment below standard pathology's detection threshold; catching them early prevents relapse
- Four primary methods cover the field: flow cytometry, PCR-based assays, NGS, and ctDNA liquid biopsy — each suited to specific cancer types and clinical scenarios
- Advanced NGS and next-generation flow can detect 1 malignant cell per 100,000–1,000,000 normal cells
- Method selection turns on cancer type, sample availability, turnaround requirements, and whether baseline tumor profiling exists
- Pre-analytical quality matters: poor cfDNA extraction directly degrades assay sensitivity and result reliability
What Is MRD?
Measurable residual disease (MRD) — formerly called minimal residual disease — describes the small population of cancer cells that remain in the body after treatment. These cells cannot be detected by standard microscopy or imaging but are identifiable through highly sensitive molecular assays capable of detecting one malignant cell in tens of thousands to hundreds of thousands of normal cells.
The field has shifted from "minimal" to "measurable" residual disease to reflect a crucial distinction: the ability to quantify residual tumor burden, not just acknowledge its presence. According to the ELN-DAVID MRD Working Party's 2025 consensus document, MRD represents "the most critical post-treatment prognostic marker for patients with acute myeloid leukemia responding to therapy by morphological criteria for complete remission."
MRD monitoring has established clinical utility in B-cell ALL, AML, CML, chronic lymphocytic leukemia (CLL), and multiple myeloma (MM), with each disease having specific thresholds that define MRD negativity:
- AML: MRD-negative is defined as <0.01% (10⁻⁴) by multiparameter flow cytometry, with molecular techniques validated to 10⁻⁴ to 10⁻⁵
- B-cell ALL: Threshold is typically 10⁻⁴ to 10⁻⁵, with PCR and NGS-based methods enabling detection at or below 10⁻⁵
- CLL: MRD negativity is defined at <10⁻⁴ (one CLL cell per 10,000 leukocytes) by four-color flow cytometry per iwCLL guidelines
- Multiple myeloma: The IMWG defines MRD negativity at a minimum sensitivity of 1 in 10⁵ nucleated cells (10⁻⁵)
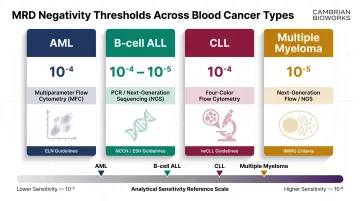
Reaching MRD-negative status at any of these thresholds signals a deeper response — and, across all four diseases, MRD result drives the next clinical decision.
Why MRD Detection Is Critical in Cancer Care
MRD status has emerged as one of the most powerful prognostic markers in modern oncology. A systematic review and meta-analysis of 81 publications encompassing 11,151 AML patients found that achievement of MRD negativity was associated with superior disease-free survival (HR 0.37) and overall survival (HR 0.36). Estimated 5-year overall survival was 68% for MRD-negative patients versus 34% for MRD-positive patients.
In multiple myeloma, pooled data from four Phase 3 trials (N=2,510) demonstrated that patients achieving complete response with MRD negativity had a progression-free survival hazard ratio of 0.20 compared to those with lesser responses or MRD-positive status. The 48-month PFS rate was 70.4% for MRD-negative versus 23.9% for MRD-positive patients.
These survival differences translate directly into treatment decisions made at every stage of care. Clinical decisions influenced by MRD results include:
- Confirms depth of remission after induction therapy
- Determines need for consolidation chemotherapy or allogeneic stem cell transplant
- Enables early intervention on molecular relapse before clinical symptoms appear
- Guides treatment de-escalation in patients with sustained MRD negativity
MRD is also reshaping clinical trial design and regulatory approval pathways. In January 2026, the FDA issued guidance recommending MRD-negative complete response as an endpoint to support accelerated approval of drugs for multiple myeloma, acknowledging its association with improved progression-free and overall survival.
The Main MRD Detection Methods Explained
Four principal MRD detection approaches are used in clinical and research settings, each differing in what they detect (cell surface markers versus genetic sequences), what sample type they require, and how deeply they can probe for residual disease. The following sections break down the technical foundations, practical requirements, and performance characteristics of each method.
Flow Cytometry and Next-Generation Flow (NGF)
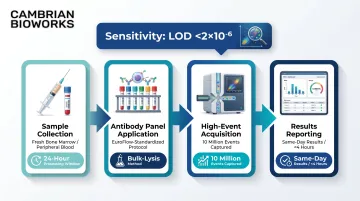
Multiparameter flow cytometry detects aberrant cell surface marker profiles (immunophenotypes) to distinguish residual malignant cells from normal hematopoietic cells. Next-generation flow (NGF) — exemplified by the EuroFlow consortium's standardized protocols — dramatically improves sensitivity over conventional flow by using optimized antibody panels, bulk-lysis procedures, and higher cell acquisition counts.
NGF achieves a validated limit of detection <2 × 10⁻⁶ and limit of quantification <5 × 10⁻⁶, based on identification of 20 or more aberrant cells among 10 million events. In fact, 25% of patients classified as MRD-negative by conventional 8-colour flow were reclassified as MRD-positive by NGF, underscoring the clinical impact of improved sensitivity.
Practical considerations:
- Fresh sample required: NGF requires fresh bone marrow or blood analyzed within 24 hours, as cell-surface antigens and light-scatter properties degrade upon freezing
- Rapid turnaround: The complete NGF procedure requires less than 4 hours, with results typically available the same day
- No prior tumor information needed: Flow cytometry can be applied to any sample without requiring baseline tumor sequencing
- Widely applicable: Particularly valuable in multiple myeloma and ALL, where aberrant immunophenotypes are well-characterized
PCR-Based Assays (RT-PCR, qPCR, Digital PCR)
PCR-based MRD detection identifies cancer-specific genetic sequences — such as fusion transcripts like BCR-ABL in CML or clonal immunoglobulin/T-cell receptor rearrangements in ALL — using highly sensitive amplification. Three formats exist:
Qualitative RT-PCR detects presence or absence of a target sequence. Quantitative PCR (qPCR) measures target abundance relative to a standard curve. Digital PCR (dPCR) partitions reactions into thousands of droplets, enabling absolute quantification without a standard curve and reducing susceptibility to amplification efficiency differences.
RT-qPCR quantification of BCR-ABL1 mRNA is the established standard for CML monitoring, with response milestones defined on the International Scale: MMR/MR³ (≤0.1%), MR⁴ (≤0.01%), MR⁴·⁵ (≤0.0032%), and MR⁵ (≤0.001%). Achieving MR⁴ requires detection sensitivity of at least 10⁻⁴, which necessitates 32,000 or more reference transcript copies in the sample.
Key characteristics:
- Requires known tumor target: PCR assays need a defined genetic sequence to amplify, established from initial diagnostic samples
- Broad sensitivity range: Typically 10⁻⁴ to 10⁻⁶ depending on assay format and input material
- Can use stored samples: Unlike flow cytometry, DNA-based PCR assays can be run on archived or frozen samples
- Widely accessible infrastructure: Most molecular diagnostics labs have the necessary equipment
Next-Generation Sequencing (NGS)
NGS-based MRD uses deep sequencing to track patient-specific clonal sequences — including immunoglobulin gene rearrangements or somatic mutations — at extremely high sensitivity. The clonoSEQ assay (Adaptive Biotechnologies) is the only FDA-cleared in vitro diagnostic for MRD monitoring in B-cell ALL, multiple myeloma, and CLL.
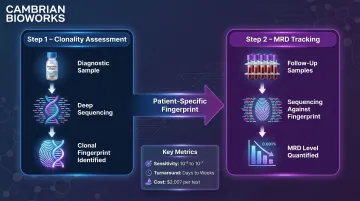
The workflow involves two steps:
- Clonality (ID) Assessment — baseline or high-disease-burden samples are sequenced to identify dominant rearranged immunoglobulin sequences specific to the malignancy
- Tracking (MRD) Assessment — subsequent samples are sequenced to detect and quantify those clonal sequences, determining MRD level over time
ClonoSEQ achieves a frequency limit of detection of 2.26 × 10⁻⁵ at 500 ng DNA input and 6.77 × 10⁻⁷ at 20 micrograms DNA input — capable of detecting one cancer cell among one million healthy cells, which is 100 times more sensitive than standard flow cytometry under optimal conditions.
Practical considerations:
- Can use stored samples: NGS is DNA-based and does not require viable cells or fresh samples
- Requires baseline tumor profile: Initial sequencing to establish clonal fingerprint is mandatory before tracking can begin
- Longer turnaround time: Centralized lab processing and batching requirements extend turnaround to days or weeks
- Higher per-test cost: Medicare Clinical Laboratory Fee Schedule rate for clonoSEQ is $2,007 USD per test as of 2025
ctDNA / Liquid Biopsy-Based MRD
Circulating tumor DNA (ctDNA) testing detects fragments of tumor-derived DNA shed into the bloodstream, enabling MRD monitoring from a peripheral blood draw rather than invasive bone marrow aspiration. This approach is the fastest-growing area of MRD testing and particularly relevant for solid tumors.
Two strategies exist:
- Tissue-informed (personalized): Requires prior tumor tissue sequencing to identify patient-specific mutations, which are then tracked in plasma with bespoke assays (e.g., Signatera, RaDaR, PhasED-Seq)
- Tissue-naive (agnostic): Uses universal panels to detect recurrent cancer-associated alterations without prior tumor sequencing (e.g., Guardant Reveal, FoundationOne Liquid CDx)
In non-small cell lung cancer (NSCLC), ctDNA clearance after neoadjuvant therapy correlates with significantly improved outcomes — the NADIM trial showed ctDNA clearance associated with PFS HR 0.16 and OS HR 0.05. In diffuse large B-cell lymphoma, end-of-treatment ctDNA positivity predicted progression with HR 13.69.
Critical pre-analytical considerations:
Genomic DNA contamination from white blood cell lysis is a major risk for ctDNA assays. Apoptotic cfDNA has a characteristic modal fragment size of approximately 166 bp, while gDNA from lysed cells consists of high-molecular-weight fragments (>10,000 bp) that dilute the target and reduce sensitivity.
Best practices to minimize contamination include:
- Use cfDNA-stabilizing collection tubes (e.g., Streck BCT, stable up to 14 days)
- Process EDTA samples within 2–6 hours of collection
- Employ two-step centrifugation to remove residual cells
- Store plasma at -80°C until extraction
- Use extraction methods specifically optimized to limit gDNA carryover
Cambrian Bioworks' Manta platform exemplifies automated cfDNA extraction designed specifically to address these pre-analytical challenges. CE-IVD certified for clinical DNA extraction, Manta completes cfDNA extraction in approximately 55 minutes with minimal genomic DNA contamination — critical for ctDNA-based MRD assays. The platform's open system compatibility allows integration with diverse assay platforms, and it has been validated for use in oncology labs including Tata Memorial Hospital and ACTREC.
Choosing the Right MRD Detection Method for Your Lab
No single MRD method is universally superior. The right choice depends on disease type, sample availability, required sensitivity, infrastructure, and turnaround time. This section provides a practical decision-making framework for oncology and molecular diagnostics labs.
Disease Type as the Primary Driver
| Cancer Type | Preferred MRD Method(s) | Guideline Source |
|---|---|---|
| Multiple Myeloma | NGF and/or NGS (10⁻⁵ threshold) | IMWG (2016 criteria) |
| CML | RT-qPCR / dPCR for BCR-ABL1 | ELN (Cross et al., 2023) |
| B-ALL | NGF and/or NGS | ELN / NCCN |
| CLL | NGS (clonoSEQ) or flow cytometry | NCCN / FDA-cleared |
| DLBCL / Lymphoma | ctDNA or NGS | Emerging evidence base |
| Solid tumors (NSCLC, CRC) | ctDNA liquid biopsy | Clinical trial data |
Sample Type and Availability
Bone marrow aspirate is required for NGF and NGS in most hematologic malignancies. Poor sample quality directly undermines MRD sensitivity — hemodilution from prolonged aspiration lowers functional sensitivity. Best practice is to assign MRD testing to a technical first pull (after needle repositioning), collecting 2–3 mL into K₂-EDTA tubes.
Peripheral blood is viable for CLL, CML BCR-ABL monitoring, and ctDNA-based MRD in solid tumors and increasingly in lymphoma. Blood draws are less invasive and support more frequent longitudinal monitoring.
Sensitivity vs. Speed vs. Cost Trade-offs
| Method | Typical Turnaround | Sensitivity | Relative Cost |
|---|---|---|---|
| NGF / Flow Cytometry | <4 hours (same day) | 10⁻⁵ to 10⁻⁶ | Low |
| RT-qPCR / dPCR | 1–3 days | 10⁻⁴ to 10⁻⁶ | Moderate |
| NGS (clonoSEQ) | Days to 2–3 weeks | 10⁻⁵ to 10⁻⁷ | High |
| ctDNA liquid biopsy | 5–14 days | Variable | Moderate to High |
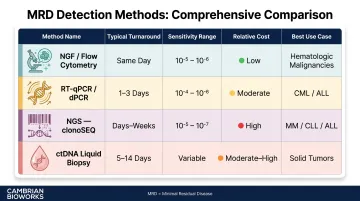
NGF offers the fastest clinical turnaround but requires fresh samples and specialized flow cytometers. NGS provides the deepest sensitivity but has the longest turnaround and highest per-test cost.
Infrastructure and Expertise Requirements
Each method carries distinct infrastructure demands:
- NGF: Multi-laser flow cytometer (e.g., BD LSRFortessa), EuroFlow-standardized panels, and dedicated analysis software (e.g., Infinicyt). Primarily available at large academic centers or referral labs.
- NGS (clonoSEQ): Runs as a centralized send-out assay. In-house NGS MRD additionally requires a sequencer, validated bioinformatics pipeline, and computational infrastructure.
- PCR/dPCR: The most broadly accessible option. Standard molecular biology equipment suffices; digital PCR platforms (Bio-Rad QX200, QIAGEN QIAcuity) are now standard in many clinical labs.
- ctDNA: Requires high-quality cfDNA extraction to minimize genomic DNA contamination. Automated sample preparation reduces variability and is essential for labs scaling ctDNA MRD programs.
How Cambrian Bioworks Supports MRD Detection Workflows
The accuracy of every downstream MRD assay — whether PCR, NGS, or ctDNA — begins with the quality of extracted nucleic acid. Cambrian Bioworks enables reliable MRD testing through automated, CE-IVD certified nucleic acid and cfDNA extraction platforms designed specifically for clinical oncology workflows.
Manta: Optimized for cfDNA-Based MRD Testing
The Manta platform addresses critical pre-analytical challenges in ctDNA-based MRD workflows:
- CE-IVD certified under Regulation (EU) 2017/746 for clinical DNA extraction
- Completes cfDNA extraction in approximately 55 minutes with no batching pressure
- Selective binding conditions limit gDNA contamination — essential for ctDNA assays where high-molecular-weight gDNA dilutes the target signal
- Delivers 5–20× higher cfDNA yield from 1 mL plasma compared to leading competitors requiring 2 mL
- Compatible with NGS-based ctDNA assays, PCR, and digital PCR workflows out of the box
- Validated at institutions including Tata Memorial Hospital and ACTREC for clinical cancer research and blood cancer monitoring

Manta's ability to process 1 to 32 samples per run — without batching pressure — makes it suitable for both high-throughput facilities and labs with variable daily sample loads.
Broader Portfolio for High-Throughput MRD Workflows
The Beluga automated liquid handler extends workflow automation beyond extraction, supporting barcode scanning, sample quantification, and direct elution into 2D barcoded tubes for full traceability. Paired with Manta, Beluga reduces manual variability and shortens turnaround time for labs running frequent MRD monitoring cycles.
Labs seeking to optimize their MRD sample preparation pipeline can connect with Cambrian Bioworks to explore platform compatibility and validation support.
Frequently Asked Questions
What are the methods of MRD detection?
The four main MRD detection methods are flow cytometry (including next-generation flow), PCR-based assays (RT-PCR, qPCR, digital PCR), next-generation sequencing (NGS), and ctDNA liquid biopsy. Each detects residual cancer cells through different molecular markers — cell surface proteins, genetic sequences, or circulating tumor DNA fragments.
What is the gold standard for diagnosing leukemia?
Bone marrow biopsy with morphological assessment has been the traditional standard. However, sensitive MRD assays (NGS and next-generation flow) now form the cornerstone of complete response evaluation in acute leukemias, offering detection sensitivity far beyond what microscopy can achieve.
Does a flow cytometry test rule out cancer?
A flow cytometry MRD-negative result means no aberrant cells were detected above the assay's sensitivity threshold, but it does not absolutely rule out cancer. Residual disease may be present below the detection limit, at other anatomic sites, or missed due to sampling error.
What is the normal range for MRD test?
MRD tests don't have a "normal range" — results are reported as MRD-negative (no detectable disease above threshold) or MRD-positive with quantification. Thresholds are disease-specific — typically 10⁻⁴ in AML and 10⁻⁵ in multiple myeloma.
What is the difference between MRD-negative and MRD-positive?
MRD-negative means no cancer cells are detectable above the assay's sensitivity threshold — an outcome associated with better long-term prognosis. MRD-positive means residual disease is still present, typically prompting additional treatment or closer monitoring.
What sample types are used for MRD testing?
Bone marrow aspirate is the standard sample for most hematologic MRD assays (next-generation flow, NGS, PCR). Peripheral blood is used for CLL, CML BCR-ABL monitoring, and ctDNA-based liquid biopsy MRD testing — particularly for solid tumors and increasingly for lymphoma.


