Measurable residual disease (MRD) testing was developed to close this detection gap. Modern MRD assays can identify as few as 1 malignant cell among 100,000 to 1 million normal cells—a 1,000- to 10,000-fold improvement in sensitivity. This article examines what MRD is, how it's measured using flow cytometry, PCR, and sequencing technologies, why it has become the strongest predictor of relapse and survival in blood cancers, and the technical and clinical challenges that remain as MRD moves from research to routine care.
TLDR
- MRD refers to cancer cells remaining after treatment that fall below the 1–5% detection threshold of standard morphology
- The "minimal" to "measurable" rename reflects that residual disease burden is quantifiable — not necessarily small — and detection levels vary by assay
- Achieving MRD-negative status correlates with dramatically better survival across ALL, AML, and other blood cancers (hazard ratios 0.23–0.37)
- Detection methods include multiparameter flow cytometry, RT-qPCR, digital PCR, and NGS, with sensitivity ranging from 10⁻³ to 10⁻⁶ depending on the assay
- Positive MRD results trigger transplant or treatment intensification; an MRD-negative result enables safe de-escalation
What Is Measurable Residual Disease?
Measurable residual disease (MRD) refers to the population of cancer cells that persist in a patient's body after treatment but remain undetectable by standard morphological assessment. Conventional microscopy and cytology can typically identify disease only when blast cells exceed approximately 1-5% of the total cell population. MRD exists in the zone below this threshold—where leukemia cells are present at levels as low as 1 per 10,000 to 1 per 1,000,000 normal white blood cells, yet still capable of causing relapse.
The term "measurable" emphasises two key concepts. First, these residual cells are not necessarily minimal in biological impact—they simply fall below the detection limit of older methods. Second, the precise level of MRD detected varies depending on which assay is used, making it essential to specify the testing method and sensitivity threshold when reporting results. The European LeukemiaNet MRD Working Party formally recommended this terminology shift in 2018 to reflect that MRD is a quantifiable biomarker rather than a qualifier of disease burden.
Where MRD Fits in the Treatment Timeline
MRD testing becomes clinically relevant after a patient achieves "complete remission" (CR) based on conventional criteria—typically defined as fewer than 5% blasts in bone marrow, normal blood counts, and absence of extramedullary disease. At this point, standard tests indicate no detectable cancer. However, MRD assays can reveal whether microscopic disease still exists, providing critical prognostic information.
MRD negativity does not equal cure. Some patients who test MRD-negative still relapse months or years later, likely due to disease below the assay's detection limit or sampling limitations.
Conversely, not all MRD-positive patients will relapse: clinical context, disease subtype, treatment phase, and the specific assay used all shape interpretation. Despite these nuances, MRD status consistently emerges as the most powerful independent predictor of relapse and survival available after treatment initiation.
Disease Contexts Where MRD Is Established
MRD testing is most clinically validated in:
- Acute lymphoblastic leukemia (ALL) — both pediatric and adult, using flow cytometry and IG/TCR gene rearrangement PCR
- Acute myeloid leukemia (AML) — primarily flow cytometry and molecular targets like NPM1 mutations
- Chronic myeloid leukemia (CML) — RT-qPCR for BCR-ABL1 transcripts is standard of care
- Chronic lymphocytic leukemia (CLL) — flow-based detection after chemoimmunotherapy or targeted therapy
- Multiple myeloma — next-generation flow and sequencing platforms for post-induction monitoring
The role of MRD is expanding into lymphomas and other hematologic malignancies, though standardisation and clinical utility remain under investigation in these settings.
How Is MRD Measured? Key Detection Methods
MRD detection relies on technologies capable of identifying rare malignant cells within a background of millions of normal cells. Four main approaches dominate current practice, each with distinct advantages, limitations, and appropriate use cases.
Multiparameter Flow Cytometry (MFC)
Multiparameter flow cytometry detects leukemia cells by identifying abnormal patterns of surface and intracellular proteins (immunophenotypes) using fluorochrome-labelled antibodies. Modern instruments use 8-12 colours simultaneously, allowing highly sensitive discrimination between normal and malignant cells.
Sensitivity: Typically 10⁻³ to 10⁻⁵ (1 in 1,000 to 1 in 100,000), with specialised protocols like EuroFlow's Next Generation Flow achieving 10⁻⁵ to 10⁻⁶ in multiple myeloma.
Two Main Strategies:
- LAIP (Leukemia-Associated ImmunoPhenotype): Identifies patient-specific aberrant markers at diagnosis, then tracks those exact markers during follow-up. Advantage: straightforward and reproducible. Limitation: susceptible to immunophenotypic shifts—cancer cells may gain or lose antigens during treatment, leading to false-negative results.
- DFN (Different from Normal): Evaluates emerging abnormal differentiation or maturation patterns at each timepoint, without strict reliance on the diagnostic phenotype. Advantage: can detect new aberrancies and monitor patients lacking diagnostic samples. Limitation: requires significant expertise and is harder to standardise across laboratories.
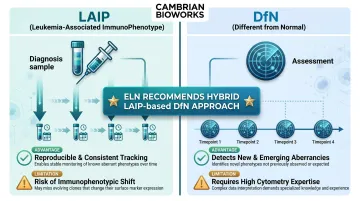
The European LeukemiaNet recommends a hybrid "LAIP-based DfN approach," combining the reproducibility of tracking known markers with the flexibility to detect phenotypic evolution.
Key Challenge: Unlike molecular methods, MFC protocols for AML lack universal standardisation. Inter-laboratory variability in antibody panels, gating strategies, and sensitivity thresholds remains a significant barrier. The ELN consensus noted that MFC approaches "have not been qualitatively or quantitatively standardised" to the same degree as PCR-based assays.
RT-qPCR (Reverse Transcription Quantitative PCR)
RT-qPCR targets specific gene rearrangements or fusion transcripts unique to the patient's leukemia, amplifying and quantifying these sequences to measure residual disease.
Sensitivity: 10⁻⁴ to 10⁻⁶, depending on assay design and input quality.
Common Targets:
- IG/TCR gene rearrangements: Present in >95% of ALL patients; standardised by the EuroMRD consortium
- BCR-ABL1: Philadelphia chromosome-positive CML and ALL
- PML-RARA: Acute promyelocytic leukemia (APL)
- NPM1 mutations: Present in ~30% of AML; the most validated mutation target for AML MRD
- CBFB::MYH11 and RUNX1::RUNX1T1: Core-binding factor AML subtypes
Standardisation Through EuroMRD: The EuroMRD consortium has developed comprehensive guidelines for IG/TCR-based RQ-PCR MRD, ensuring identical interpretation across laboratories. The consortium conducts regular quality assurance rounds with inter-laboratory comparisons, achieving high reproducibility for ALL monitoring.
Requirement for Diagnostic Material: PCR-based MRD requires identification of patient-specific targets from a diagnostic (pretreatment) sample. At least two distinct IG/TCR rearrangements are recommended to reduce false-negative risk from clonal evolution. This dependency can limit applicability when diagnostic material is unavailable or insufficient.
Digital PCR (dPCR)
Digital PCR partitions each sample into thousands of individual droplets or microwells, performing PCR in each compartment independently. After amplification, the system counts positive versus negative partitions to provide absolute quantification without requiring standard curves.
Advantages Over RT-qPCR:
- Absolute quantification without calibrators or reference standards
- Reduced sensitivity to PCR inhibitors due to compartmentalisation
- Higher precision for rare variants (detects down to 0.001% mutant allele frequency)
- In CML monitoring, ddPCR detected BCR-ABL1 positivity three months earlier than qPCR in some patients, and identified residual disease in 24 of 32 samples that tested negative by conventional qPCR
Current Status: While technically promising, standardised clinical guidelines for dPCR MRD assays are still under development. Most labs continue to rely on validated RT-qPCR protocols, particularly where regulatory and reimbursement frameworks favour established methods.
Next-Generation Sequencing (NGS)
NGS-based MRD uses high-throughput sequencing to identify and quantify clonal IG/TCR gene rearrangements, somatic mutations, or other leukemia-specific sequences.
Sensitivity: 10⁻⁵ to 10⁻⁶ (frequency-based) or even lower with error-corrected sequencing technologies.
Key Advantages:
- Does not require patient-specific primer design for follow-up samples (unlike traditional PCR)
- Can detect clonal evolution—identifies new subclones emerging during treatment
- Enables comprehensive mutation profiling across multiple genes simultaneously
- ClonoSEQ, the first FDA-cleared NGS-based MRD assay, achieved a limit of detection of 6.77 × 10⁻⁷ at 20 micrograms DNA input for B-cell ALL and multiple myeloma
Limitations:
- Requires bioinformatics expertise and computational infrastructure
- Higher sequencing error rates historically limited sensitivity, though duplex UMI (unique molecular identifier)-based error correction is addressing this
- Costlier per sample than flow cytometry or PCR in many settings
- Provisional rather than definitive MRD thresholds in AML (ELN 2022 suggests ≥0.1% variant allele frequency, excluding clonal hematopoiesis mutations)
The Critical Role of Nucleic Acid Quality
All PCR and NGS-based MRD methods depend on high-quality DNA or RNA input. Poor-quality or contaminated nucleic acids extracted from bone marrow or blood can produce false-negative results, unreliable quantification, or complete assay failure—directly compromising clinical decision-making.
Key quality parameters include:
- DNA integrity: High molecular weight DNA with minimal fragmentation ensures efficient amplification
- Yield thresholds: Adequate input quantity (typically 10-20 micrograms for sensitive NGS assays) is essential
- Purity ratios: A260/A280 values of 1.8-2.0 and A260/A230 values of 2.0-2.2 indicate minimal protein or chemical contamination
- Absence of PCR inhibitors: Residual proteins, haem, or extraction reagents can block amplification
Cambrian Bioworks' Manta is a CE-IVD and CDSCO-certified automated DNA extraction system that uses magnetic bead-based purification to meet these quality requirements. It processes 1-32 samples in under 30 minutes and delivers A260/280 ratios of 1.8-2.0 with high molecular weight DNA—reducing per-test costs by up to 56% compared to manual workflows.
For oncology and molecular diagnostics labs scaling MRD testing, that combination of throughput, consistency, and validated output quality is what separates reliable assay results from avoidable pre-analytical failures.
Clinical Significance of MRD in Hematologic Malignancies
MRD status after treatment has emerged as the single most powerful predictor of relapse and survival across hematologic malignancies, consistently outperforming traditional risk factors in multivariate analyses.
Prognostic Impact: Meta-Analytic Evidence
Acute Lymphoblastic Leukemia (ALL):
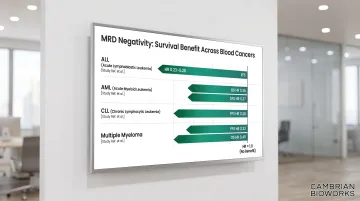
A 2017 meta-analysis (Berry et al., JAMA Oncology) pooled 39 studies including 13,637 patients with pediatric and adult ALL. Patients achieving MRD negativity demonstrated:
- Pediatric EFS: Hazard ratio 0.23 (95% BCI 0.18-0.28)—a 77% reduction in risk of relapse or death
- Adult EFS: Hazard ratio 0.28 (95% BCI 0.24-0.33)
- Overall survival: Hazard ratios of 0.28 for both pediatric and adult cohorts
These results held across MFC-based, PCR-based, and fusion transcript detection methods.
The same pattern holds in myeloid disease. Acute Myeloid Leukemia (AML):
Short et al. (JAMA Oncology 2020) analyzed 81 publications encompassing 11,151 AML patients and found:
- Disease-free survival: HR 0.37 (95% CrI 0.34-0.40)
- Overall survival: HR 0.36 (95% CrI 0.33-0.39)
The association remained significant across MFC, molecular (PCR), and NGS subgroups. Cytogenetics/FISH did not show significant MRD association (OS HR 0.77), underscoring that assay sensitivity matters.
Other Malignancies:
- Chronic lymphocytic leukemia: PFS HR 0.28 (95% CI 0.20-0.39)—a 72% reduction in progression or death
- Multiple myeloma: PFS HR 0.33 (95% CI 0.29-0.37); OS HR 0.45 (95% CI 0.39-0.51)
MRD-Guided Therapy Escalation
Patients with persistent or high-level MRD after initial therapy face substantially elevated relapse risk. Treatment protocols increasingly use MRD to identify candidates for intensified intervention:
- Earlier referral to allogeneic stem cell transplant (allo-SCT) in first remission, before genetics-based staging alone would indicate
- Blinatumomab for MRD-positive B-ALL — the first therapy with an FDA-approved MRD-specific indication (full approval June 2023) for patients in CR1/CR2 with MRD ≥0.1%
- Addition of maintenance therapy or alternative regimens when MRD converts from negative to positive
MRD-Enabled Therapy De-Escalation
Conversely, early MRD negativity identifies patients who may safely receive reduced treatment intensity, sparing them from chemotherapy-related toxicity without compromising cure rates.
The Dutch Childhood Oncology Group ALL10 study demonstrated that chemotherapy was substantially reduced in one-quarter of children with ALL selected on undetectable MRD, with outcomes comparable to standard-intensity arms.
This principle—escalating for MRD-positive patients, reducing burden for MRD-negative patients—defines the current model of risk-adapted oncology.
Surveillance and Early Relapse Detection
MRD monitoring during remission can detect molecular relapse weeks to months before clinical or morphologic relapse becomes evident. This lead time is particularly well-established in:
- Acute promyelocytic leukemia (APL): RT-qPCR monitoring of PML-RARA detects molecular relapse a median of 3 months ahead of hematological relapse. Pre-emptive treatment at molecular relapse yields 2-year PFS of 85%, versus 44% when delayed
- Core-binding factor AML: RUNX1::RUNX1T1 and CBFB::MYH11 transcript monitoring enables early intervention
Guidelines recommend RT-qPCR every 3 months for the first 3 years after APL consolidation, exemplifying how MRD has transitioned from research tool to standard-of-care surveillance.
MRD in Clinical Practice: From Risk Stratification to Drug Development
Integration Into Treatment Algorithms
Different diseases have different guideline-endorsed timepoints for MRD assessment:
- ALL: End of induction (day 29-33), end of consolidation, pre-transplant
- AML: Post-induction, pre-consolidation, post-consolidation, pre-transplant
- CML: Every 3 months indefinitely per ELN 2020 guidelines
In chronic myeloid leukemia, molecular monitoring has achieved the deepest integration. ELN 2020 recommendations mandate RT-qPCR BCR-ABL1 measurement at defined intervals, with specific response milestones triggering treatment decisions:
- Molecular response milestones: MMR/MR3 (BCR-ABL1 ≤0.1%), MR4 (≤0.01%), MR4.5 (≤0.0032%)
- Treatment failure thresholds: BCR-ABL1 >10% at 3 or 6 months, or >1% at 12 months prompts TKI switch
- Discontinuation criteria: Requires CML-CP, >5 years TKI, sustained deep molecular response (≥MR4) for >2 years, with monthly monitoring for 6 months post-cessation
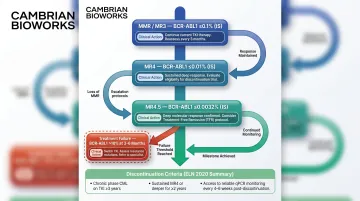
CML's structured framework—where each assay result maps to a defined clinical action—shows what MRD-guided care looks like when the evidence base is mature. Regulatory agencies have taken notice, and are now examining whether this model can accelerate drug approval in other haematologic malignancies.
MRD as a Clinical Trial Surrogate Endpoint
Because MRD negativity correlates strongly with survival across meta-analyses, regulatory agencies are exploring MRD as an accelerated approval endpoint to expedite drug development. The FDA has issued guidance recognising MRD as a potential surrogate endpoint specifically for multiple myeloma, and granted De Novo clearance to ClonoSEQ (Adaptive Biotechnologies) for B-cell ALL and multiple myeloma MRD monitoring from bone marrow DNA.
Adoption has been uneven, though. An FDA analysis found nearly 40% of proposals to incorporate MRD into prescribing information were rejected — reflecting persistent questions about assay standardisation, optimal thresholds, and the strength of the MRD-survival correlation in some disease contexts. Until these gaps close, MRD's role as a regulatory endpoint will remain disease-specific rather than universal.
Global Adoption Disparity
Academic centres in high-income countries have integrated MRD into standard care, but community practices and laboratories in low- and middle-income countries face significant barriers:
- Infrastructure gaps: Limited access to flow cytometers, PCR platforms, or sequencing capacity
- Cost: High consumable costs, reagent dependencies, and reimbursement uncertainty
- Standardisation: Lack of harmonised protocols and quality assurance programmes outside consortium networks
This disparity creates a widening outcome gap. Patients in resource-constrained settings who could benefit most from MRD-guided therapy de-escalation often lack access to the testing needed to act on it. For laboratories in India and similar markets, this is where accessible, validated nucleic acid extraction tools become operationally critical — enabling reliable sample preparation as the foundation for any downstream MRD assay. Cambrian Bioworks' automated extraction platforms are built specifically for this context: compact, CE-IVD certified, and designed to fit the throughput and cost realities of clinical labs scaling up molecular diagnostics.
Challenges and Limitations in MRD Testing
Despite its proven clinical value, MRD testing faces technical, biological, and implementation challenges that limit consistent adoption and interpretation across labs.
Technical Limitations and Standardisation
Three persistent technical issues complicate MRD measurement in practice:
- Inter-laboratory variability: MFC-based MRD for AML remains heterogeneous — labs differ in antibody panels, gating strategies, cell acquisition numbers, and sensitivity thresholds. An inter-laboratory comparison study identified numerous false-positive results. Flow-based approaches have not been standardised to the degree achieved for IG/TCR PCR through EuroMRD.
- Specimen quality and haemodilution: Bone marrow aspirate is the preferred specimen, and the ELN strongly recommends submitting the first pull for MRD analysis. Later pulls suffer from peripheral blood contamination (haemodilution), evidenced by >90% mature neutrophils, producing falsely low MRD results. Peripheral blood MRD is generally an order of magnitude less sensitive.
- Method-dependent thresholds: A patient may be MRD-negative by one assay (10⁻⁴ sensitivity MFC) but MRD-positive by another (10⁻⁶ NGS). Clinical decisions must account for which method was used and its validated threshold.

Biological Complexity of Interpretation
Even a technically valid MRD result requires careful biological interpretation:
- Clonal haematopoiesis (DTA) mutations: Mutations in DNMT3A, TET2, and ASXL1 are common in older adults and can persist after AML treatment without indicating residual malignancy. The ELN 2022 AML guidelines state: "Mutations consistent with premalignant clonal haematopoiesis (e.g., DNMT3A, TET2, ASXL1) should not be considered as MRD." Excluding these targets is essential to avoid false positives and unnecessary treatment escalation.
- MRD positivity ≠ imminent relapse: Not all MRD-positive patients relapse. Disease kinetics (rising vs. stable vs. declining), MRD level, treatment phase, and disease biology all influence outcome.
- MRD negativity ≠ cure: Some MRD-negative patients still relapse — likely due to disease below the detection limit, spatial sampling gaps (aspiration covers only one site), or sanctuary sites not reached by the assay.
Emerging Solutions and Future Directions
Several developments are actively addressing these gaps:
- Liquid biopsy (ctDNA): Technologies like CAPP-seq and PhasED-seq can detect circulating tumour DNA at frequencies around 0.002%, offering less invasive sampling and broader anatomical coverage than bone marrow aspiration. Clinical validation for haematologic MRD is still early-stage — NCCN currently gives ctDNA MRD only a category 2B recommendation in PET-positive DLBCL when biopsy is not feasible.
- AI-assisted analysis: Machine learning algorithms are being developed to improve MFC gating consistency and identify subtle leukaemic populations in heterogeneous samples.
- Multi-omic profiling: Combining mutation tracking, immunophenotyping, and epigenetic signatures can improve sensitivity and reduce false negatives from single-platform approaches.
- Global harmonisation: EuroMRD and EuroFlow consortia continue to develop standardised protocols and conduct international quality assurance rounds to reduce variability and expand MRD testing capacity.
Frequently Asked Questions
What is measurable residual disease?
Measurable residual disease (MRD) refers to the small population of cancer cells remaining in the body after treatment that are undetectable by standard microscopy or cytology. Modern molecular and flow-based assays can detect these cells at sensitivities as low as 1 in 100,000 to 1 in 1,000,000.
What is the new name for minimal residual disease?
The preferred term is now "measurable residual disease" (still abbreviated MRD). The change reflects that biological impact isn't always minimal, and detection levels vary by testing method.
How to measure measurable residual disease?
MRD is measured using four main methods: multiparameter flow cytometry (MFC), RT-qPCR for gene rearrangements or fusion transcripts, digital PCR (dPCR), and next-generation sequencing (NGS). Method choice depends on disease type, available molecular targets, and required sensitivity.
How to interpret MRD results?
MRD-negative results indicate lower relapse risk and better prognosis; MRD-positive results suggest residual disease and may prompt therapy escalation. Interpretation must account for:
- The specific assay and its detection threshold
- Disease subtype and treatment phase
- Broader clinical context
Are you ever cured from AML?
Long-term remission and functional cure are possible in AML, particularly with allogeneic stem cell transplant in high-risk patients. Relapse remains a risk even years after treatment, but sustained MRD negativity is associated with the best long-term outcomes.
Can you have leukemia for years and not know it?
Some leukemias — chronic lymphocytic leukemia (CLL) and low-grade myelodysplastic syndromes (MDS) — can be present for years before symptoms emerge. MRD testing detects residual disease after treatment in diagnosed patients; it is not a general population screening tool.


