CamSelect Long
CamSelect Long
CamSelect Long
CamSelect Long
CamSelect Long
Increase usable Gb per long-read sequencing flow cell by up to 3x
Increase usable Gb per long-read sequencing flow cell by up to 3x
Increase usable Gb per long-read sequencing flow cell by up to 3x
Increase usable Gb per long-read sequencing flow cell by up to 3x

The flow cell
efficiency problem
Long-read sequencing platforms have finite capacity per run, constrained by pore availability and functional lifetime. Short DNA fragments waste sequencing capacity by occupying pores unproductively, accelerating pore attrition and limiting achievable read lengths.
CamSelect Long addresses this challenge through tunable cutoff (under 2kb, 5kb, 7kb and 10kb), enriching libraries for longer fragments.
This improves the proportion of informative long reads, increases read N50 and pass yield, stabilizes run performance, and enables greater usable data output per flow cell.
The flow cell
efficiency problem
Long-read sequencing platforms have finite capacity per run, constrained by pore availability and functional lifetime. Short DNA fragments waste sequencing capacity by occupying pores unproductively, accelerating pore attrition and limiting achievable read lengths.
CamSelect Long addresses this challenge through tunable cutoff (under 2kb, 5kb, 7kb and 10kb), enriching libraries for longer fragments.
This improves the proportion of informative long reads, increases read N50 and pass yield, stabilizes run performance, and enables greater usable data output per flow cell.
The flow cell
efficiency problem
Long-read sequencing platforms have finite capacity per run, constrained by pore availability and functional lifetime. Short DNA fragments waste sequencing capacity by occupying pores unproductively, accelerating pore attrition and limiting achievable read lengths.
CamSelect Long addresses this challenge through tunable cutoff (under 2kb, 5kb, 7kb and 10kb), enriching libraries for longer fragments.
This improves the proportion of informative long reads, increases read N50 and pass yield, stabilizes run performance, and enables greater usable data output per flow cell.
The flow cell
efficiency problem
Long-read sequencing platforms have finite capacity per run, constrained by pore availability and functional lifetime. Short DNA fragments waste sequencing capacity by occupying pores unproductively, accelerating pore attrition and limiting achievable read lengths.
CamSelect Long addresses this challenge through tunable cutoff (under 2kb, 5kb, 7kb and 10kb), enriching libraries for longer fragments.
This improves the proportion of informative long reads, increases read N50 and pass yield, stabilizes run performance, and enables greater usable data output per flow cell.
A new standard for long read preparation
A new standard for long read preparation
A new standard for long read preparation
A new standard for long read preparation
Improves read N50 and long-read fraction
Increased usable Gb per flowcell for long read sequencing
Increases pass yield (Q ≥ threshold)
Increases pass yield (Q ≥ threshold)
Enhances pore retention and flow-cell efficiency
Low input requirement (≈1 µg DNA)
Low input requirement (≈1 µg DNA)
Automation-ready magnetic bead chemistry
Automation-ready magnetic bead chemistry
Purpose-built for long-read sequencing economics
Purpose-built for long-read sequencing economics
Purpose-built for long-read sequencing economics
Purpose-built for long-read sequencing economics
Maximize data yield per read
Maximize data yield per read
Maximize data yield per read
Maximize data yield per read
CamSelect Long improves data yield and N50 while reducing pore consumption, delivering more contiguous assemblies.
Libraries were prepared using the ONT Native Barcoding Ligation kit (EXP-NBD114) according to manufacturer’s protocol, and sequenced on a PromethION 2 Solo platform with FLO-PRO114M flowcell to a target depth of 5 Gb per sample.
CamSelect Long improves data yield and N50 while reducing pore consumption, delivering more contiguous assemblies.
Libraries were prepared using the ONT Native Barcoding Ligation kit (EXP-NBD114) according to manufacturer’s protocol, and sequenced on a PromethION 2 Solo platform with FLO-PRO114M flowcell to a target depth of 5 Gb per sample.
CamSelect Long improves data yield and N50 while reducing pore consumption, delivering more contiguous assemblies.
Libraries were prepared using the ONT Native Barcoding Ligation kit (EXP-NBD114) according to manufacturer’s protocol, and sequenced on a PromethION 2 Solo platform with FLO-PRO114M flowcell to a target depth of 5 Gb per sample.
CamSelect Long improves data yield and N50 while reducing pore consumption, delivering more contiguous assemblies.
Libraries were prepared using the ONT Native Barcoding Ligation kit (EXP-NBD114) according to manufacturer’s protocol, and sequenced on a PromethION 2 Solo platform with FLO-PRO114M flowcell to a target depth of 5 Gb per sample.
Metric
Metric
No cleanup
(control)
No cleanup
(control)
CamSelect
Long
CamSelect
Long
Improvement
(%)
Improvement
(%)
Metric
No cleanup
(control)
CamSelect
Long
Improvement
(%)
Metric
No cleanup
(control)
CamSelect
Long
Improvement
(%)
Metric
No cleanup
(control)
CamSelect
Long
Improvement
(%)
Total yield
1.7 Gb
5.5 Gb
223.50%
Total yield
1.7 Gb
5.5 Gb
223.50%
Read length N50
4,745 bp
7419 bp
56.40%
Read length N50
4,745 bp
7419 bp
56.40%
Mean read length
3416 bp
5152 bp
50.80%
Mean read length
3416 bp
5152 bp
50.80%
Long read fraction>10kb
4.70%
11.90%
153.20%
Long read fraction>10kb
4.70%
11.90%
153.20%
Stool-derived and saliva-derived DNA was analyzed on the Agilent TapeStation System before and after treatment with CamSelect LongTM. The results are depicted below.
Stool-derived and saliva-derived DNA was analyzed on the Agilent TapeStation System before and after treatment with CamSelect LongTM. The results are depicted below.
Stool-derived and saliva-derived DNA was analyzed on the Agilent TapeStation System before and after treatment with CamSelect LongTM. The results are depicted below.
Stool-derived and saliva-derived DNA was analyzed on the Agilent TapeStation System before and after treatment with CamSelect LongTM. The results are depicted below.
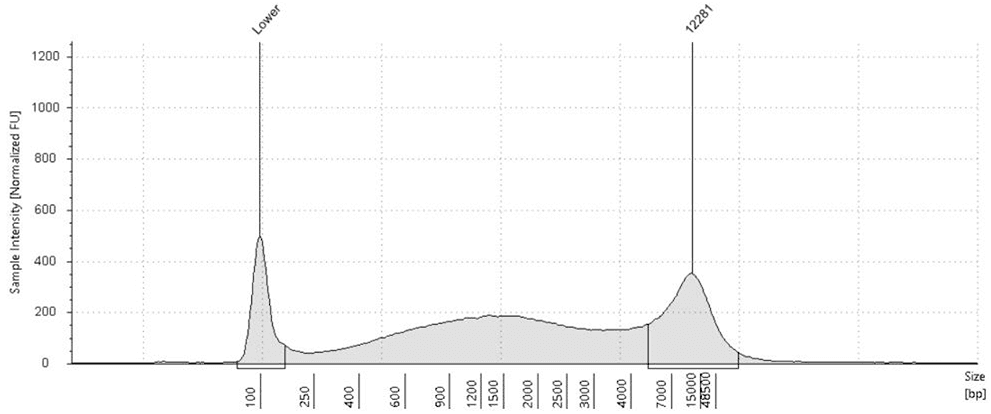
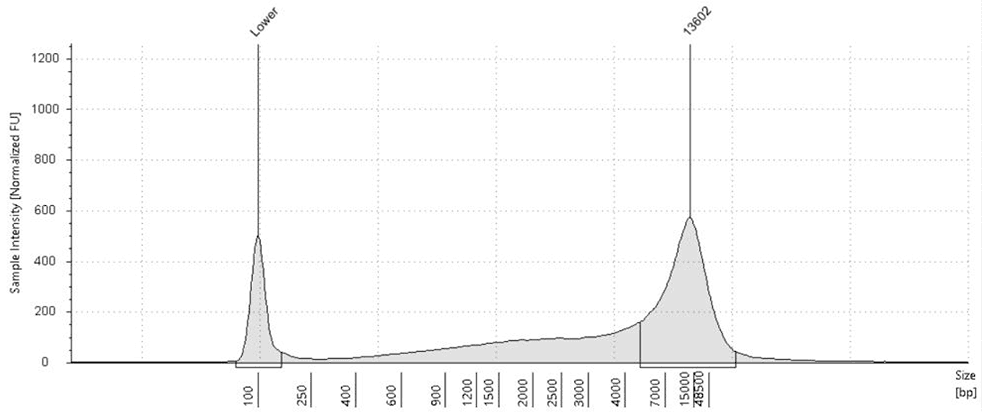
Fig.1: TapeStation analysis shows Stool DNA size selection pre-treatment (left) vs post-treatment (right) with CamSelect Long". Cleans up complex stool-derived DNA by reducing short fragments <4 kb.
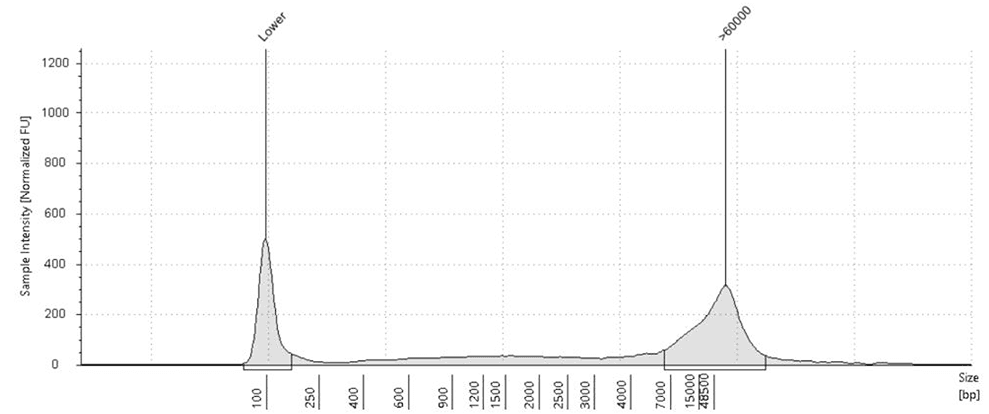
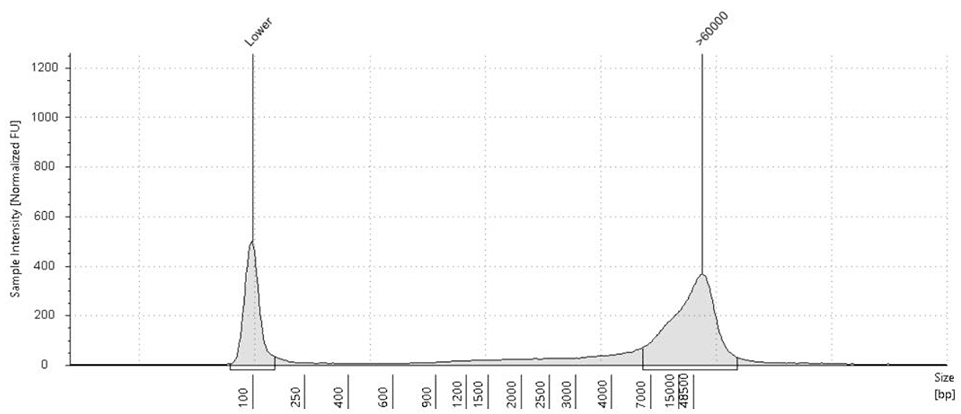
Fig.2: Saliva-derived DNA before (left) vs after (right) treatment with CamSelect Long show selective depletion of short fragments (<7 kb) & enrichment of HMW DNA.


Fig.1: TapeStation analysis shows Stool DNA size selection pre-treatment (left) vs post-treatment (right) with CamSelect Long". Cleans up complex stool-derived DNA by reducing short fragments <4 kb.


Fig.2: Saliva-derived DNA before (left) vs after (right) treatment with CamSelect Long show selective depletion of short fragments (<7 kb) & enrichment of HMW DNA.


Fig.1: TapeStation analysis shows Stool DNA size selection pre-treatment (left) vs post-treatment (right) with CamSelect Long". Cleans up complex stool-derived DNA by reducing short fragments <4 kb.


Fig.2: Saliva-derived DNA before (left) vs after (right) treatment with CamSelect Long show selective depletion of short fragments (<7 kb) & enrichment of HMW DNA.


Fig.1: TapeStation analysis shows Stool DNA size selection pre-treatment (left) vs post-treatment (right) with CamSelect Long". Cleans up complex stool-derived DNA by reducing short fragments <4 kb.


Fig.2: Saliva-derived DNA before (left) vs after (right) treatment with CamSelect Long show selective depletion of short fragments (<7 kb) & enrichment of HMW DNA.
Post analysis
Post analysis
Untreated
sample
Untreated
sample
Sample treated with
CamSelect Long
Sample treated with
CamSelect Long
Post analysis
Untreated
sample
Sample treated with
CamSelect Long
Post analysis
Untreated
sample
Sample treated with
CamSelect Long
Post analysis
Untreated
sample
Sample treated with
CamSelect Long
% reads mapping to human host
1.17
0.278
% reads mapping to human host
1.17
0.278
Assembled data
79 mb
192 mb
Assembled data
79 mb
192 mb
MAGS
43
116
MAGS
43
116
Read integrity and contiguity
Read integrity and contiguity
Read integrity and contiguity
Read integrity and contiguity
In stool samples, cleanup and size-selection prior to library preparation reduced host-mapped reads, increased total assembled sequence length, and substantially improved recovery of metagenome-assembled genomes (MAGs).
In stool samples, cleanup and size-selection prior to library preparation reduced host-mapped reads, increased total assembled sequence length, and substantially improved recovery of metagenome-assembled genomes (MAGs).
In stool samples, cleanup and size-selection prior to library preparation reduced host-mapped reads, increased total assembled sequence length, and substantially improved recovery of metagenome-assembled genomes (MAGs).
In stool samples, cleanup and size-selection prior to library preparation reduced host-mapped reads, increased total assembled sequence length, and substantially improved recovery of metagenome-assembled genomes (MAGs).
Workflow overview
Workflow overview
Workflow overview
Workflow overview
Workflow overview
CamSelect Long enables rapid size-selective enrichment of high-molecular-weight DNA for long-read sequencing. Optimized magnetic bead chemistry retains fragments above a defined cutoff while removing shorter DNA, yielding purified DNA compatible with downstream library preparation.
CamSelect Long enables rapid size-selective enrichment of high-molecular-weight DNA for long-read sequencing. Optimized magnetic bead chemistry retains fragments above a defined cutoff while removing shorter DNA, yielding purified DNA compatible with downstream library preparation.
CamSelect Long enables rapid size-selective enrichment of high-molecular-weight DNA for long-read sequencing. Optimized magnetic bead chemistry retains fragments above a defined cutoff while removing shorter DNA, yielding purified DNA compatible with downstream library preparation.
CamSelect Long enables rapid size-selective enrichment of high-molecular-weight DNA for long-read sequencing. Optimized magnetic bead chemistry retains fragments above a defined cutoff while removing shorter DNA, yielding purified DNA compatible with downstream library preparation.
Add reagents
Add reagents
Add reagents
Add reagents
Incubate
PCR amplification
PCR amplification
Incubate
Magnetize
Magnetize
Magnetize
Magnetize
Wash & Airdry
Wash & Airdry
Wash & Airdry
Wash & Airdry
Elute
Elute
Elute
Elute
QC
QC
QC
QC
Technical specifications
Technical specifications
Technical specifications
FAQs
FAQs
FAQs
How CamSelect Long compares
How CamSelect Long compares
How CamSelect Long compares
How CamSelect Long compares
The only automation-ready bead-based solution with no instrument required
The only automation-ready bead-based solution with no instrument required
The only automation-ready bead-based solution with no instrument required
The only automation-ready bead-based solution with no instrument required
Product
Product
Pack Size
Pack Size
Price
Price
Cost/Prep
Cost/Prep
Method
Method
Automation
Automation
Notes
Notes
Product
Pack Size
Price
Cost/Prep
Method
Automation
Notes
Product
Pack Size
Price
Cost/Prep
Method
Automation
Notes
CamSelect Long
CamSelect Long
Cambrian
Cambrian
25 preps
25 preps
$500
$500
$20
$20
Bead
Bead
Yes
Yes
No instrument required
No instrument required
CamSelect Long
Cambrian
25 preps
$500
$20
Bead
Yes
No instrument required
CamSelect Long
Cambrian
25 preps
$500
$20
Bead
Yes
No instrument required
Circulomics SRE Kit
Circulomics SRE Kit
PacBio/Sage
PacBio/Sage
6 preps
6 preps
$299
$299
$50
$50
Manual
Manual
No
No
Targets <10–25 kb fragments
Targets <10–25 kb fragments
Circulomics SRE Kit
PacBio/Sage
6 preps
$299
$50
Manual
No
Targets <10–25 kb fragments
Circulomics SRE Kit
PacBio/Sage
6 preps
$299
$50
Manual
No
Targets <10–25 kb fragments
BluePippin Cassettes
BluePippin Cassettes
Sage Science
Sage Science
10 cassettes
10 cassettes
$625-725
$625-725
$12-18
$12-18
Gel
Gel
No
No
Requires BluePippin instrument
Requires BluePippin instrument
BluePippin Cassettes
Sage Science
10 cassettes
$625-725
$12-18
Gel
No
Requires BluePippin instrument
BluePippin Cassettes
Sage Science
10 cassettes
$625-725
$12-18
Gel
No
Requires BluePippin instrument
ONT SFE Kit
ONT SFE Kit
Oxford Nanopore
Oxford Nanopore
30 reactions
30 reactions
$120
$120
$4
$4
Manual
Manual
No
No
Requires ≥3 µg input DNA
Requires ≥3 µg input DNA
ONT SFE Kit
Oxford Nanopore
30 reactions
$120
$4
Manual
No
Requires ≥3 µg input DNA
ONT SFE Kit
Oxford Nanopore
30 reactions
$120
$4
Manual
No
Requires ≥3 µg input DNA
Improve pore utilization
Improve pore utilization
Improve pore utilization
Improve pore utilization
Get more usable genomic data from every sequencing run.
Get more usable genomic data from every sequencing run.
Get more usable genomic data from every sequencing run.
Get more usable genomic data from every sequencing run.