Introduction
Transplant recipients face a lifelong challenge: the risk of organ rejection. Current monitoring relies heavily on tissue biopsies—invasive procedures that carry procedural risks, delay timely intervention, and generate costs averaging thousands of dollars. In heart transplants, endomyocardial biopsy carries a 1.6% complication rate, yet surveillance biopsies detect treated rejection in only 1.2% of cases. For the 173,727 organs transplanted globally in 2024, this creates both clinical and economic pressure to find better alternatives.
Donor-derived cell-free DNA (dd-cfDNA) circulating in the recipient's bloodstream addresses this directly. When donor organ cells undergo injury or death from rejection, infection, or other causes, they release DNA fragments into the blood. Droplet digital PCR (ddPCR) detects and quantifies these fragments with limits of detection as low as 0.002% of total circulating DNA.
This article examines how ddPCR-based dd-cfDNA testing works, why it outperforms alternatives for routine monitoring, what clinical thresholds apply across organ types, and the pre-analytical steps labs must follow for reliable results.
TLDR:
- dd-cfDNA testing detects organ injury earlier than traditional biopsies by measuring donor DNA in recipient blood
- ddPCR provides absolute quantification with 0.002% sensitivity—far superior to qPCR for low-fraction detection
- Validated thresholds: 1.0% for kidney rejection, 0.20% for heart, with high negative predictive value
- Pre-analytical quality—especially minimising gDNA contamination—directly impacts accuracy
- CE-IVD certified automated extraction platforms ensure consistent longitudinal results
What Is Donor-Derived Cell-Free DNA and Why Does It Matter?
dd-cfDNA consists of short DNA fragments (typically 150–200 base pairs) released from dying or damaged donor organ cells into the recipient's bloodstream. These fragments circulate alongside the recipient's own cell-free DNA. The detection challenge is real: donor DNA typically represents less than 1% of total cfDNA in stable transplant recipients, dropping below 0.5% between rejection episodes.
Clinical Relevance for Transplant Monitoring
Rising dd-cfDNA levels correlate strongly with active rejection episodes, including both antibody-mediated rejection (ABMR) and T cell-mediated rejection (TCMR). The DART study established the foundational 1.0% threshold for kidney transplant rejection, reporting 59% sensitivity, 85% specificity, and a 96% negative predictive value for ABMR. This high NPV demonstrates dd-cfDNA's primary clinical value: confidently ruling out rejection and avoiding unnecessary biopsies.
Beyond rejection detection, dd-cfDNA levels can:
- Rise as early as 5 months before biopsy-confirmed rejection in heart transplants
- Track whether anti-rejection therapy is working through serial measurements
- Detect subclinical rejection before functional decline appears in conventional biomarkers
Organ-specific baseline and rejection thresholds:
| Organ | Stable Baseline | Rejection Threshold | Key Study |
|---|---|---|---|
| Kidney | 0.19% median | ≥1.0% | Bloom/DART 2017 |
| Heart | 0.06–0.6% | 0.20% | Khush 2019 |
| Lung | 0.05–0.06% | 1.0% | De Vlaminck 2015 |
| Liver | 2.2–3.3% | 11.2% | Schutz 2017 |
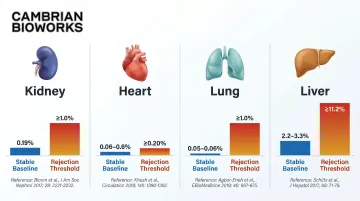
These thresholds reflect biological differences: smaller organs like the heart shed less DNA into a larger blood volume and therefore need lower detection thresholds, while larger organs produce higher baseline fractions.
The Measurement Challenge
Accurately quantifying dd-cfDNA means detecting a signal that may represent less than 0.5% of total cfDNA, a fraction well below the reliable detection range of conventional qPCR. This drives the need for absolute quantification methods that can:
- Maintain precision at low template concentrations
- Resist amplification efficiency variation between runs
- Eliminate reliance on standard curves, which introduce error at low copy numbers
How Droplet Digital PCR Works for dd-cfDNA Detection
Droplet digital PCR fundamentally changes how we count DNA molecules. Instead of measuring fluorescence accumulation during amplification (as qPCR does), ddPCR partitions each 20 µL reaction into up to 20,000 nanolitre-sized water-in-oil droplets. Each droplet acts as an independent micro-reaction containing either zero or one target DNA molecule.
Core Quantification Principle
After thermal cycling (typically 40 cycles), droplets are read individually for fluorescence. Droplets containing target DNA fluoresce positively; empty ones remain dark. Poisson statistics applied to the ratio of positive-to-negative droplets yield absolute target concentration, with no calibration curves or reference standards required.
The result is exceptional quantification precision. Clausen et al. (2023) validated 40 SNP assays on the Bio-Rad QX200 platform and achieved:
- Limit of detection: 3 copies per reaction (0.002% fraction)
- Limit of quantification: 35 copies per reaction (0.038% fraction, CV <25%)
- Precision at 0.5% fraction: CV of 9.1–13.6%
- Linearity: Pearson r = 0.997 for absolute values
Determining Donor Fraction Using Informative Alleles
ddPCR determines donor fraction through SNP-based or HLA-based target strategies. Labs identify genetic loci where donor and recipient are homozygous for different alleles (so-called "informative SNPs"), which occur at approximately 12.5% of tested loci when both alleles carry ~50% population frequency.
The workflow:
- Genotype both donor and recipient pre-transplant to identify informative SNPs
- Design dual-probe assays targeting donor-specific and recipient-specific alleles
- Run both targets in the same reaction (multiplexed)
- Calculate donor fraction: (donor-positive droplets) / (total positive droplets)
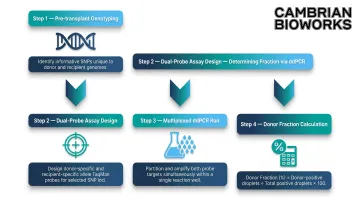
Only 8 µL cfDNA input per 20 µL reaction is required, and multiple SNPs can be queried simultaneously in a single well, increasing informativeness without additional sample volume.
Why Absolute Quantification Matters
Transplant monitoring requires tracking small percentage changes over time: a shift from 0.3% to 1.2% dd-cfDNA can signal active rejection. ddPCR's ability to provide both fractional (%) and absolute (copies/mL) quantification offers meaningful clinical advantages over fraction-only reporting.
Oellerich et al. (2019) demonstrated that absolute dd-cfDNA quantification achieved an AUC-ROC of 83% versus 73% for fractional determination in kidney rejection detection. Bohmer et al. (2025) confirmed this advantage in heart transplant, where absolute dd-cfDNA at 25 copies/mL achieved AUC 0.87 compared to 0.75 for donor fraction alone.
Recipient-derived cfDNA fluctuates with leukocyte counts, immunosuppression changes, and intercurrent illness. These fluctuations alter the denominator in fractional calculations independently of graft injury, potentially masking or exaggerating the true donor signal. Absolute quantification bypasses this confounding variable entirely.
Advantage Over qPCR for Rare-Target Detection
qPCR relies on Ct-value thresholds and standard curves, both of which introduce substantial error at low template concentrations. When donor DNA represents <0.5% of total cfDNA, Ct variability makes reliable quantification difficult. ddPCR's binary counting approach is inherently resistant to:
- PCR inhibitors (they affect all droplets equally)
- Amplification efficiency variation between runs
- Template quality differences
- Low DNA concentration effects
For labs tracking patients monthly or quarterly over years, that robustness translates directly to consistent, comparable results across longitudinal monitoring.
Clinical Applications of ddPCR-Based dd-cfDNA Testing in Transplant Medicine
Kidney Transplant Monitoring
Kidney transplant represents the best-studied application of dd-cfDNA testing. The DART trial (Bloom et al. 2017) established the 1.0% threshold for active rejection (TCMR >1B or ABMR), reporting 61% positive predictive value and 84% negative predictive value. Subsequent studies refined these findings:
- Huang et al. (2019): Strong ABMR discrimination (AUC 0.82) but not TCMR (AUC 0.42) — pointing to organ- and rejection-type-specific performance
- Sigdel et al. (2018): SNP-based multiplex PCR achieved 88.7% sensitivity and 72.6% specificity at 1.0% threshold (NPV 95.1%)
- Dauber et al. (2020): INDEL PCR identified a 2.7% threshold for TCMR (AUC 0.84)
The biological explanation: ABMR causes more endothelial damage and cell death than pure TCMR, releasing more donor DNA into circulation. This makes dd-cfDNA most informative for detecting antibody-mediated injury.
Heart and Lung Transplant Surveillance
In cardiothoracic transplantation, repeated biopsies carry higher procedural risk than in kidney. Endomyocardial biopsy complications occur in 1.6% of procedures, with odds ratios of 12.74 for biopsies performed within the first month post-transplant. ddPCR-based dd-cfDNA testing enables more frequent, lower-risk surveillance.
Heart transplant evidence:
- Khush et al. (2019) established 0.2% threshold with 97.1% NPV in 740 patients
- Bohmer et al. (2025) demonstrated absolute dd-cfDNA at 25 copies/mL achieved AUC 0.87 for symptomatic rejection
- De Vlaminck et al. (2014) showed dd-cfDNA elevation up to 5 months before biopsy-proven rejection
Lung transplant evidence:
- Novo et al. (2024) found median dd-cfDNA of 0.222% during CLAD versus 0.051% in stable patients (AUC 0.709-0.780)
- Donor fraction was not elevated during infectious complications, suggesting it may help discriminate CLAD from infection
- Jang et al. (2021) reported median dd-cfDNA of 1.95% during acute rejection with 77% sensitivity and 84% specificity at 1.0% threshold
Post-Rejection Treatment Monitoring
Serial dd-cfDNA measurements track whether anti-rejection therapy is effective. Declining donor fractions over days to weeks indicate treatment response, providing clinicians a quantitative tool beyond serum creatinine or clinical symptoms. This application depends entirely on ddPCR's precision at low allele frequencies — detecting a clinically meaningful drop from 1.5% to 0.4% requires:
- Consistent droplet partitioning across runs
- Reproducible absolute quantification without standard curves
- Low coefficient of variation at sub-1% allele frequencies
Early Subclinical Rejection Detection
The ADMIRAL study (Bu et al. 2022) demonstrated that dd-cfDNA at a 0.5% cutoff can identify subclinical kidney rejection—biopsy-confirmed rejection without clinical symptoms or functional decline. In liver transplant, dd-cfDNA levels rose nearly 2 weeks before biopsy confirmation. That 2-week lead time represents a concrete clinical window — enough time to adjust immunosuppression before graft function deteriorates.
Limitations and Clinical Context
dd-cfDNA can also be elevated in non-rejection events, including:
- Viral infections (CMV, BKV)
- Ischemia-reperfusion injury
- Surgical trauma
- Cholestasis/steatohepatitis (liver)
- Mixed chimerism
- Pregnancy (foetal cfDNA interference)
Clinical interpretation must combine dd-cfDNA data with patient history, immunosuppression levels, DSA status, and conventional biomarkers. ISHLT's December 2023 position statement captures the current consensus well:
"High negative predictive value, allowing clinicians to safely rule out active rejection" — while emphasizing that elevated values still require biopsy confirmation.
ddPCR vs. Other Methods for dd-cfDNA Quantification
ddPCR vs. Next-Generation Sequencing
NGS-based approaches (AlloSure, Prospera) analyse hundreds to thousands of SNPs simultaneously without requiring prior donor genotyping—a significant operational advantage for deceased-donor transplants where donor tissue may be unavailable.
Key trade-offs:
| Parameter | ddPCR | NGS |
|---|---|---|
| Turnaround time | ~1 working day | 2-3 working days |
| Cost per test | ~$400 (₹33,000) | ~$2,700 (₹2,25,000) |
| LOD (fraction) | 0.002% | Platform-dependent |
| Quantification | Absolute (cp/mL) + fraction (%) | Fraction (%) typically |
| Prior genotyping | Required | Optional (genotyping-free methods available) |
| Infrastructure | Bio-Rad QX200/QX600 | Sequencer + bioinformatics pipeline |
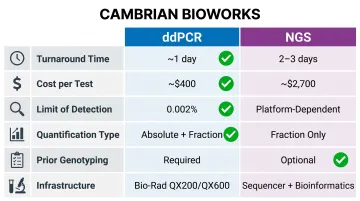
These economics become clearer in longitudinal programmes. Once the upfront genotyping investment is made, ddPCR enables cost-effective in-house testing—and for patients monitored monthly or quarterly over years, the per-test economics strongly favour it.
NGS is better suited for initial assessment, genome-wide research applications, or settings where donor tissue is unavailable. Its key limitation: most NGS platforms report only fractional dd-cfDNA, forgoing the absolute quantification that clinical validation studies have consistently supported.
ddPCR vs. Conventional qPCR
qPCR lacks the precision for low-fraction detection required in dd-cfDNA testing. When donor fractions fall below 0.5%, Ct variability makes reliable quantification difficult. qPCR's reliance on standard curves and threshold cycle values introduces systematic errors at low template concentrations, whereas ddPCR's binary counting approach maintains consistent precision across the entire dynamic range.
Decision Framework
- Choose ddPCR for: Routine clinical surveillance with pre-identified informative SNPs, cost-sensitive programmes, labs requiring fast turnaround, and applications where absolute quantification adds clinical value
- Choose NGS for: Initial genotyping-free assessment, research requiring genome-wide resolution, and settings where donor tissue is unavailable for pre-transplant genotyping
Sample Preparation: The Critical First Step for Accurate dd-cfDNA Testing
cfDNA extraction quality directly determines ddPCR accuracy. Low yield, co-purification of high-molecular-weight genomic DNA from white blood cells, or degradation during processing can distort the apparent dd-cfDNA fraction and generate false results.
Importance of Minimising gDNA Contamination
Genomic DNA contamination is the primary pre-analytical confounding factor. If leukocyte-derived high-MW gDNA from the recipient is co-extracted, it dilutes the true cfDNA pool and artificially lowers the apparent dd-cfDNA fraction, potentially masking active rejection.
Best practices include:
- Gentle centrifugation protocols (first spin at 800-1,600 × g for 10 minutes, then high-speed second spin at 16,000 × g for 10 minutes)
- Prompt plasma separation (process EDTA tubes within 4-6 hours)
- Extraction methods that enrich short cfDNA fragments (150-200 bp)
- Selective binding conditions that limit high-MW gDNA capture
Shin et al. (2022) demonstrated that delayed processing of EDTA tubes beyond 4-6 hours causes a "remarkable increase" in total DNA from white blood cell lysis, diluting low-concentration dd-cfDNA signal and potentially causing false-negative results.
Automated Extraction for Consistency
Manual extraction introduces inter-operator variability that creates inconsistency in dd-cfDNA fraction measurements across time points. When tracking small changes over weeks or months, a shift from 0.4% to 0.9% must reflect true biological change, not processing noise.
Automated, magnetic bead-based platforms standardise workflow and reduce that variability. Platforms like Cambrian Bioworks' Manta, which carries CE-IVD certification for clinical DNA extraction, achieve tighter reproducibility than manual spin-column methods.
The Manta system processes 1-32 samples per run with cfDNA extraction completed in approximately 27-30 minutes, using pre-filled cartridges that minimise hands-on time and contamination risk.

Automated extraction using magnetic bead technology can achieve:
- Minimises gDNA contamination through selective binding conditions
- Maximises cfDNA yield from limited plasma volumes (5-20× higher from 1 mL vs. competitor methods requiring 2 mL)
- Maintains consistent fragment size enrichment (150-200 bp)
- Elution into barcoded tubes for seamless ddPCR integration
Sample Collection and Storage Recommendations
Collection tubes and hold times:
- EDTA tubes (K2/K3): Process within 4-6 hours at room temperature or 4°C
- Streck Cell-Free DNA BCT: Stable for 14 days at 6-37°C
- PAXgene Blood ccfDNA Tubes: Stable for 7 days at 15-25°C
- Roche Cell-Free DNA Collection Tube: Stable for 7 days at 18-25°C
Centrifugation protocol:
- First spin: 800-1,600 × g for 10 minutes (separate plasma from cellular components)
- Second spin: 16,000 × g for 10 minutes (remove residual leukocytes) — for Streck tubes, use 300 × g for 20 min followed by 5,000 × g for 10 min instead
Storage and freeze-thaw:
- Separated plasma: 4°C for up to 3 hours short-term
- Long-term: -20°C or -80°C with only one freeze-thaw cycle permitted
- Always aliquot plasma to avoid repeated freeze-thaw
- Quantify cfDNA using fluorometry (Qubit) or automated electrophoresis (TapeStation), not spectrophotometry

Frequently Asked Questions
What is a normal dd-cfDNA level after a successful transplant?
Baseline dd-cfDNA in stable recipients is typically very low: kidney recipients show median 0.19%, heart recipients range from 0.06-0.6%, and lung recipients show median 0.05-0.06%. Levels vary by organ type and time post-transplant, with transient elevation immediately post-surgery typically resolving within 7-14 days.
How does ddPCR differ from NGS for dd-cfDNA testing?
ddPCR offers absolute, targeted quantification at roughly 7× lower cost and faster turnaround (1 day vs. 2-3 days) using pre-selected informative SNPs. NGS provides genome-wide resolution without prior donor genotyping but demands greater infrastructure and bioinformatics capacity.
Can dd-cfDNA testing replace biopsy for rejection diagnosis?
No. dd-cfDNA testing is currently used as a non-invasive complement to biopsy rather than a replacement, helping guide when biopsy is warranted and reducing unnecessary procedures. High NPV (96-99%) enables confident rule-out of rejection, but confirmatory biopsy remains standard in most clinical protocols when dd-cfDNA is elevated.
What sample type is used for dd-cfDNA testing by ddPCR?
Plasma separated from whole blood by centrifugation is the standard sample type; cfDNA-stabilising collection tubes (Streck, PAXgene, Roche) are recommended if processing is delayed beyond 4-6 hours. Minimum input is typically 1 mL plasma, yielding 100-1000 ng cfDNA sufficient for ddPCR workflows.
How sensitive is ddPCR for detecting low fractions of dd-cfDNA?
ddPCR can reliably detect and quantify allele fractions as low as 0.002% (limit of detection) and 0.038% (limit of quantification), making it sensitive enough to detect early subclinical rejection when donor cfDNA levels are still well below the 0.2-1.0% clinical thresholds used across organ types.
What factors other than rejection can elevate dd-cfDNA levels?
Viral infections (CMV, BKV), ischemia-reperfusion injury, surgical trauma, and cholestasis in liver transplant can all transiently elevate dd-cfDNA. Because these non-immunological causes are indistinguishable from rejection by dd-cfDNA alone, clinical context and conventional biomarkers are essential for interpretation.