Introduction
DNA extraction is the rate-limiting step in high-throughput PCR workflows. Poor extraction quality directly impacts amplification reliability, leading to failed tests, repeat runs, and wasted resources in both clinical and research lab settings. A stark illustration: extraction method choice can shift PCR detection rates from 8.2% to 61.2% on identical stool specimens, according to a 2022 comparative study published in Diagnostics. The difference wasn't yield — it was purity. The high-performing method delivered lower raw DNA concentration but superior inhibitor clearance, enabling reliable amplification.
That purity problem compounds at scale. Modern molecular diagnostic labs, biobanks, and sequencing facilities routinely process hundreds to thousands of samples per day. Manual extraction at this volume introduces operator-dependent variability, increases turnaround times, and creates a resource bottleneck that no team can sustain.
Choosing the right extraction method determines whether your PCR workflow succeeds or stalls.
This guide helps molecular biology and diagnostic labs match extraction method to throughput, sample type, and PCR quality requirements — so every decision is grounded in your actual workflow constraints, not just reagent cost.
TLDR
- Extraction methods range from crude chemical lysis (fast, low-cost) to automated magnetic bead systems (high purity, scalable)
- Extraction method directly determines PCR success — purity matters more than yield
- Open automated platforms offer chemistry flexibility; closed cartridge systems deliver validated walk-away ease
- Check A260/A280 (~1.8–2.0) and A260/A230 (>1.8); always run spiked controls to catch PCR inhibition early
- In clinical labs, CE-IVD certification and ISO 13485 compliance are non-negotiable alongside technical performance
Why High-Throughput DNA Extraction Matters for PCR
The Scale Challenge
The global PCR and real-time PCR testing market reached approximately $10.1 billion in 2023 (Yahoo Finance, 2024), reflecting sustained post-pandemic demand for molecular diagnostics. Meanwhile, the automated nucleic acid extraction market was valued at $4.31 billion in 2025 and is projected to reach $11.21 billion by 2035 (SNS Insider, 2025).
Molecular diagnostic labs, AMR surveillance programs, biobanks, and sequencing facilities now routinely process hundreds to thousands of samples daily. Manual extraction at this volume simply doesn't scale.
For 15 samples alone, manual extraction consumes 66 minutes of hands-on time versus 19 minutes automated — and that gap compounds with every batch. Operator-dependent variability and processing bottlenecks translate directly into delayed clinical results.
When Extraction Quality Fails, So Does the PCR
Residual inhibitors in extracted DNA cause false negatives, non-specific amplification, and polymerase inhibition. The source depends on sample type — common culprits include:
- Heme from blood (binds nucleic acids and enzymes)
- Humic acids from stool or soil samples
- Polysaccharides from plant tissue (co-precipitate with DNA)
- Residual chaotropic salts from silica column buffers (denature polymerase)
- Paraffin and formalin from FFPE tissue processing

A 2019 study in PLOS One analyzing 3,152 clinical samples found PCR inhibition in 2.0% of samples overall, with sputum and placenta particularly prone. At scale, even a 2% inhibition rate across thousands of daily samples means dozens of actionable results delayed or missed per day.
Application Diversity Demands Adaptable Systems
Different applications process different matrices at different scales:
- Infectious disease testing — blood, swabs, viral transport media
- Oncology mutation panels — FFPE tissue, cfDNA from plasma
- AMR gene detection — microbial cultures, stool, environmental samples
- Prenatal screening — whole blood, amniotic fluid
- Pharmacogenomics — buccal swabs, saliva
Each sample type carries unique inhibitor profiles and requires extraction systems that can adapt without sacrificing quality.
Core High-Throughput DNA Extraction Methods Explained
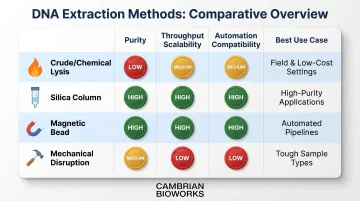
All DNA extraction methods involve five fundamental steps: cell lysis, lysate clearing, DNA binding, washing, and elution. The difference lies in purity, throughput scalability, and automation compatibility.
Crude and Chemical Lysis Methods
Methods like HotSHOT (boiling samples in dilute NaOH followed by Tris neutralization) and alkaline lysis rapidly disrupt cells and release DNA without full purification. A 2025 study in Plant Methods demonstrated 96-well crude extraction completing in 3–4 hours at £0.11/sample versus £3.68 for commercial kits (costs reported in GBP by the study authors).
Where it excels:
- Extremely fast (<15 minutes) with very low cost per sample
- Sufficient template for short-amplicon PCR
- Well-suited to 96-well genotyping or screening workflows
Where it falls short:
- Retains variable inhibitor loads depending on sample matrix
- Not recommended for NGS, digital PCR, or sensitive clinical diagnostics
- Plant polysaccharides, blood heme, and seed lipids carry over — performance depends heavily on dilution strategy and PCR master mix inhibitor tolerance
Column and Silica Membrane-Based Methods
DNA binds to silica membranes under high-chaotropic salt conditions. Contaminants wash away with alcohol-salt buffers, and purified DNA elutes in low-salt buffer — producing reliable A260/A280 ratios suited to most downstream PCR applications.
The format has a ceiling, though. While 96-well silica plates exist, they require vacuum manifolds and manual plate handling. That hands-on requirement limits true walk-away automation and reduces scalability compared to magnet-based systems.
Magnetic Bead-Based Methods
DNA binds to functionalized paramagnetic beads in chaotropic conditions. A magnet captures beads through sequential washing steps, and DNA is thermally eluted. Because magnets replace centrifugation, the format works directly with robotic liquid handlers — which is why it has become the backbone of clinical diagnostic automation.
An SPRI bead study in BMC Genomics (2023) documented 97.70% recovery efficiency for fragments 150 bp to 10 kbp, equivalent to commercial AMPure XP beads. That performance profile explains broad adoption:
- Best combination of purity, scalability, and reproducibility
- Validated across whole blood, FFPE, cfDNA plasma, swabs, and microbial cultures
- Compatible with 96-well and 384-well formats
That automation compatibility translates directly into instrument design. Cambrian Bioworks' Manta processes 1–32 samples in 27–30 minutes using magnetic bead chemistry, fits inside a biosafety hood, and delivers A260/A280 of 1.8–2.0 and A260/A230 of 2.0–2.2.
Mechanical and Physical Disruption Methods
Bead beating and tissue homogenization are essential for samples that chemical lysis alone cannot crack: fungal spores, plant tissue, and gram-positive bacteria. A 2019 study in Scientific Reports illustrated the gap — gram-positive cells and fungi remained "almost intact" after 10 minutes of bead beating that destroyed gram-negative cells entirely.
In high-throughput workflows, physical disruption is a pretreatment step rather than a standalone method. It precedes chemical or bead-based extraction to ensure complete cell lysis before the purification stage begins. Newer 96-well mechanical disruption devices are emerging, but most labs still apply this upstream rather than as a primary extraction platform.
Manual vs. Automated: Navigating Your Options
Hands-On Time and Reproducibility
A 2008 comparison study in the Journal of Molecular Diagnostics processing 15 samples showed:
| Method | Hands-On Time | Cost per Sample | Recovery vs. Manual |
|---|---|---|---|
| Manual (QIAamp) | 66 min 33 sec | $6.41 | 100% (reference) |
| easyMAG (automated) | 19 min 26 sec | $9.57–$12.95 | 86–107% |
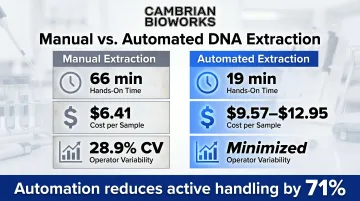
Automated systems reduce active handling by 71% while processing equivalent sample numbers with measurably higher consistency.
Manual extraction introduces substantial variability: plasma DNA extraction efficiencies showed a coefficient of variation of 28.9% across specimens (O'Connell et al., Lab Med, 2017). Automation minimises this operator-to-operator variance.

Open vs. Closed System Trade-Offs
Open platforms (such as Hamilton STAR or Tecan):
- Adaptable to many protocols and extraction chemistries
- Holmberg et al., JoVE, 2013 demonstrated 96 blood samples processed in under 1 hour with A260/A280 of 1.96
- Flexibility for labs working with diverse sample types
- Higher validation burden (user-defined protocols)
Closed cartridge-based systems:
- Purpose-built for specific applications
- Pre-validated protocols with pre-filled reagents
- Minimal training required
- True walk-away processing
- Less customisation
The Batch-Size Problem
Many conventional automated systems require near-full 96-well plates to run efficiently, creating queuing delays for lower-volume diagnostic labs. Compact, flexible automated systems that process any batch size — from a single sample to a full plate — eliminate this bottleneck and support on-demand processing for urgent clinical samples.
Cambrian Bioworks' Manta system processes 1–32 samples per run at consistent yield, using pre-filled cartridges that eliminate reagent waste when working with smaller batches.
Regulatory-Relevant Reproducibility
Clinical labs operating under ISO 13485 or EU IVDR must document lot-to-lot and operator-to-operator consistency. Automated systems with pre-filled cartridges support this by:
- Minimising reagent variability across runs
- Generating audit-ready run logs automatically
- Reducing human-introduced deviation in documented workflows
How Sample Type Affects Your Extraction Strategy
Key Sample Categories and Extraction Challenges
Whole blood and saliva:
- Common inhibitors include heme, IgG, and lactoferrin
- Thorough magnetic bead washing or blood-validated lysis buffers clear these effectively
- Typical yield: 6–15 ng/µL with A260/A280 of 1.8–2.0
FFPE tissue:
- Formalin-induced cross-linking and fragmentation require deparaffinisation and reversal steps before extraction
- Yields are low with short fragments (~200 bp), but sufficient for targeted PCR panels
- Critical threshold: A 2023 study in Pathology International identified DIN ≥2.1 as the minimum for successful genomic profiling — 0% success was recorded below this cutoff
cfDNA from plasma:
- Extremely low input (1–10 ng/mL) with fragment size ~160–200 bp demands high-sensitivity extraction
- Must preserve short fragments without co-purifying excess genomic DNA
- Cambrian Bioworks' OncoSeek cfDNA Extraction Kit delivers 5–20× higher cfDNA yield from 1 mL plasma compared to competitors requiring 2 mL
Microbial and environmental samples:
- Diverse cell wall compositions and mixed inhibitor profiles require mechanical pre-lysis combined with chemical extraction
- Standard bead beating alone may be insufficient for fungi and gram-positive bacteria
Application-Specific Kit Validation Matters
A method optimised for whole blood PBMC DNA may over-shear cfDNA fragments or under-recover DNA from fibrous FFPE tissue. Most modern automated platforms address this with application-specific kits — dedicated cfDNA plasma kits, for instance, behave very differently from genomic DNA blood kits. Using the wrong kit is one of the most common and preventable causes of PCR failure.
Key validation parameters to confirm before kit selection:
- Sensitivity and recovery rates for the target sample type
- Inhibitor clearance data for known matrix-specific inhibitors
- Fragment size preservation (critical for cfDNA and FFPE workflows)
This is especially consequential in regulated settings. Under EU IVDR or CDSCO, extraction systems must carry documented performance validation data — covering sensitivity, specificity, and inhibitor clearance — for each specific sample type in use.
Key Quality Metrics Before Running PCR
Spectrophotometric Purity Ratios
| Ratio | Target | Concern Threshold | Contaminant |
|---|---|---|---|
| A260/A280 | ~1.8–2.0 | Below 1.6 | Protein, phenol |
| A260/A230 | 2.0–2.2 | Below 1.5 | Salts, carbohydrates, phenol |
Values below target indicate inhibitors that suppress Taq polymerase. Values above 2.0 for A260/A280 may indicate RNA carryover.
When Fluorometric Quantification is Essential
For low-input samples (cfDNA, FFPE, forensic), spectrophotometry overestimates DNA concentration by 3–4 times compared to Qubit fluorometry. NanoDrop's lower detection limit is 2 ng/µL versus 0.005 ng/µL for Qubit HS. For cfDNA below 1.35 ng/µL, NanoDrop is "neither precise nor accurate" (Khetan et al., Asian J Transfus Sci, 2019).
Use fluorometry for:
- cfDNA from plasma
- FFPE samples
- Forensic samples
- Any concentration below 2 ng/µL
DNA Integrity Assessment
| Application | Integrity Requirement | Assessment Method |
|---|---|---|
| Standard PCR (amplicons <500 bp) | Moderate fragmentation tolerable | 1% agarose gel or TapeStation/Bioanalyzer |
| Longer amplicons or NGS | DIN ≥7 | Microfluidic analysis (TapeStation/Bioanalyzer) |
FFPE samples typically show lower DIN and require adjusted PCR conditions.
Inhibitor Detection Using Spiked Controls
Spectrophotometry cannot detect non-absorbing inhibitors like heme, humic acids, or polysaccharides. A spiked control addresses this gap directly.
To detect inhibition:
- Add a defined internal control (synthetic template or co-extracted spike) to the PCR reaction
- Run the reaction under standard conditions
- Measure recovery of the spiked control
- Flag suppressed recovery as inhibitor-related failure

This approach is increasingly required in clinical molecular diagnostic SOPs (Hoorfar et al., J Clin Microbiol, 2003).
DNA Concentration Optimization for PCR
| Application | Recommended Input |
|---|---|
| Standard PCR (plasmid template, 50 µL) | 0.1–1 ng |
| Standard PCR (genomic DNA, 50 µL) | 5–50 ng |
| qPCR | 0.1–100 ng (target-dependent) |
| Maximum per reaction | No more than 1 µg |
Elution buffer considerations: Eluting in low-TE (10 mM Tris-HCl, 0.1 mM EDTA) or nuclease-free water is preferred. Standard TE with 1 mM EDTA can chelate Mg²⁺ required by Taq polymerase if excess DNA is carried over.
How to Choose the Right High-Throughput Extraction Platform
Three Primary Decision Axes
Flexibility shapes which platform fits your lab's daily reality:
- Labs with diverse sample types benefit from open automated platforms supporting multiple chemistries
- Labs with standardised inputs can rely on closed systems with pre-validated protocols
Daily throughput requirements vary more than most procurement checklists acknowledge:
- Validate whether the system processes variable batch sizes or requires minimum plate fill
- Traditional batch-centric systems require near-full plates, creating delays in lower-volume labs
- Newer compact extractors handle 1–32 samples without efficiency loss
Downstream application determines how stringent your purity requirements are:
- PCR genotyping and screening tolerates crude or column-purified DNA
- NGS, digital PCR, and clinical IVD assays require fully purified, inhibitor-free DNA from validated automated systems
Regulatory Compliance in Clinical Labs
For extraction systems used in regulated IVD workflows:
EU IVDR (Regulation 2017/746) requires:
- Analytical sensitivity, specificity, trueness, and precision documentation
- Repeatability and reproducibility data
- Software lifecycle documentation for automated systems
ISO 13485 quality management:
- Documented audit trails for all device-related processes
- Lot-to-lot consistency verification
A system meeting both frameworks will carry CE-IVD marking alongside ISO 13485 certification. Cambrian Bioworks' Manta, for instance, holds both — fitting inside a biosafety hood, processing 1–32 samples without queuing pressure, and using pre-filled magnetic bead cartridges to minimise reagent variability.
Practical Evaluation Questions
Before selecting a platform, ask:
- Can it process the sample volume range we work with?
- Is it validated for our specific sample types?
- What is the walk-away time per run?
- Does it integrate with our LIMS or barcoding system?
- Is the system supported by IVD certification for regulated tests?
- What is the cost-per-sample including consumables and service?
Frequently Asked Questions
What is the difference between crude DNA extraction and purified DNA extraction for PCR?
Crude extraction releases DNA rapidly without removing inhibitors — sufficient for short-amplicon PCR when paired with inhibitor-tolerant reagents. Purified extraction uses silica or magnetic bead chemistry to remove proteins, salts, and contaminants, producing high-quality DNA required for NGS, digital PCR, or clinical diagnostics.
How much DNA is typically needed for high-throughput PCR?
Standard endpoint PCR requires 1–10 ng template DNA per reaction. qPCR works from 0.1–100 ng depending on target abundance. cfDNA and rare variant applications may require pre-amplification from lower inputs. Check concentration by fluorometry before high-throughput PCR runs.
What are the most common PCR inhibitors introduced during DNA extraction?
Heme (from blood), humic acids (from stool/soil), polysaccharides (from plants/FFPE), residual chaotropic salts (from silica column buffers), and paraffin/organic solvents (from FFPE processing). Detect inhibitors using spiked internal controls and remove through thorough washing steps in magnetic bead-based extraction.
How do I choose between column-based and magnetic bead-based extraction for high-throughput workflows?
Column-based methods provide high purity but require centrifugation or vacuum, limiting full automation. Magnetic bead-based systems allow magnet-controlled washing without centrifugation, enabling seamless robotic integration and consistent 96-well processing. For true high-throughput automated workflows, magnetic bead-based systems are the preferred choice.
Can automated DNA extraction systems handle small or variable batch sizes efficiently?
Older batch-centric systems require near-full plates for efficiency, creating delays. Newer compact automated extractors — such as the Manta by Cambrian Bioworks — handle any batch size without efficiency loss, from a single urgent sample to a full 96-well plate. This makes them well-suited for clinical diagnostic settings where daily volumes fluctuate.
What quality checks should I run before using extracted DNA in PCR?
Run the following checks before PCR:
- Purity ratios: A260/A280 ~1.8–2.0; A260/A230 >1.8 by spectrophotometry
- Concentration: Fluorometry for low-input or cfDNA samples
- Integrity: Gel or microfluidic analysis when amplicon length exceeds 300 bp
- Inhibition check: Spiked internal control PCR run for complex sample matrices


