Introduction
More than one billion formalin-fixed paraffin-embedded (FFPE) tissue samples are currently archived in hospitals and biobanks worldwide, representing the single largest molecular resource in clinical oncology and retrospective research. For rare tumors and archival studies, FFPE blocks are often the only material available. Yet the very chemistry that makes long-term preservation possible — formalin fixation — also creates a fundamental analytical barrier: covalent crosslinks between proteins and nucleic acids that severely degrade molecular yield and quality.
Standard high-temperature extraction (90°C for 90 minutes) achieves only 88% of fresh-frozen proteome coverage, leaving a persistent 10–15% gap driven by residual formaldehyde crosslinks. Those same crosslinks compromise nucleic acid recovery: FFPE sequencing libraries show approximately 50% lower true unique coverage than fresh-frozen samples, with fragment sizes often too short for reliable variant calling.
This blog explores how chemical decrosslinking — the use of small-molecule compounds to reverse formaldehyde-induced crosslinks — is closing this gap. We'll examine recent compound evaluations (including 3,4-diaminobenzoic acid as the top-performing agent), how these methods integrate into spatial proteomics and clinical NGS workflows, and what the findings mean for oncology labs, histopathology centers, and molecular diagnostics facilities working with archival tissue.
TL;DR
- FFPE fixation preserves tissue morphology but introduces crosslinks that cut proteome coverage by 10–15% and drop sequencing library quality by ~50%
- Chemical decrosslinking uses catalytic reagents to reverse these crosslinks beyond what heat alone achieves
- 3,4-diaminobenzoic acid (3,4-DABA) increased protein identifications by 24% at 70°C in recent spatial proteomics research
- Recovery of cysteine-containing peptides improves significantly, surfacing biologically relevant proteins missed by standard protocols
- Applications span spatial proteomics, cancer biomarker discovery, archival NGS, and clinical FFPE diagnostics
What Is Chemical Decrosslinking in FFPE Tissues?
Chemical decrosslinking is the use of specific small-molecule compounds to catalytically reverse formaldehyde-induced covalent crosslinks in FFPE tissue. Formalin reacts with nucleophilic amino acid residues — lysine, cysteine, histidine, arginine, tryptophan, and tyrosine — to form three primary modification types:
| Modification | Mass Shift | Stability | Mechanism |
|---|---|---|---|
| Methylol adduct | +30 Da | Reversible (heat-labile) | Hydroxymethyl group addition |
| Schiff base | +12 Da | Moderately reversible | Dehydration of methylol adduct |
| Methylene bridge | +12 Da | Stable; difficult to reverse | Covalent crosslink between residues |
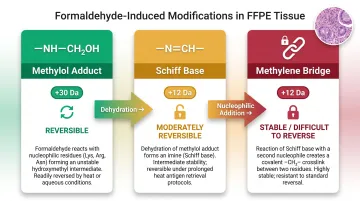
These crosslinks create both intra-molecular (within a single protein or nucleic acid strand) and inter-molecular (protein-to-protein, protein-to-DNA, DNA-to-RNA) bonds that standard extraction methods cannot fully eliminate.
Contrast with Thermal Decrosslinking
Standard high-temperature FFPE extraction (90°C, 90 minutes) partially reverses heat-labile methylol adducts but leaves stable methylene bridges largely intact. Research quantifying this gap found that FFPE tissues achieve only 88% of fresh-frozen proteome coverage under optimized thermal conditions — a 10–15% shortfall attributed specifically to loss of hydrophilic peptides carrying unresolved crosslinks.
Thermal methods also introduce their own limitations for nucleic acid recovery:
- High temperatures can denature double-stranded DNA, reducing downstream amplification efficiency
- Further fragmentation of already-damaged RNA creates a direct trade-off between crosslink reversal and additional degradation
- Methylene bridges — the most stable modification type — remain largely unaffected regardless of treatment duration
Origins in RNA/DNA Chemistry
The conceptual foundation for chemical decrosslinking comes from Karmakar et al.'s 2015 work in Nature Chemistry. Their study demonstrated that water-soluble bifunctional organocatalysts (specifically anthranilates and phosphanilates) accelerate formaldehyde adduct reversal on nucleic acid bases. These catalysts increased amplifiable RNA yields from FFPE samples by 7–25-fold compared to standard commercial extraction kits, at mild temperatures of 37–55°C.
The organocatalytic logic first validated for nucleic acids has since been extended to protein decrosslinking in mass spectrometry proteomics workflows — testing compounds that share structural features with the original catalysts to address the same methylene bridge problem at the protein level.
How Chemical Decrosslinking Works in FFPE Sample Preparation
Chemical decrosslinking is not a standalone step — it's embedded within a multi-stage FFPE workflow where reagent choices, temperature, and timing all shape final molecular yield. Understanding where and how it fits reveals why so many standard protocols leave performance on the table.
Step 1 – Deparaffinization
Paraffin wax embedding must be removed before molecular extraction can proceed. Traditional methods use xylene followed by ethanol rehydration, but non-toxic alternatives have emerged:
- Hot distilled water (90–95°C for 15 minutes): comparable protein yield to xylene, eliminates toxic solvents
- Heptane/methanol or limonene-based solutions: no impact on downstream RNA quality
- Commercial deparaffinization kits: compatible with automated workflows
While deparaffinization method has minimal impact on nucleic acid quality, solvent selection matters for lab safety — and directly affects which downstream steps are compatible.
Step 2 – Proteolytic Digestion and Protein Extraction
Proteinase K incubation lyses tissue and brings crosslinked proteins and nucleic acids into solution. In nanoPOTS-LCM workflows (Nanodroplet Processing in One pot for Trace Samples coupled with laser capture microdissection), sample processing occurs at nanoliter scale — where buffer composition becomes especially consequential.
Extraction buffer composition is where chemical decrosslinking agents are introduced. A typical nanoPOTS buffer contains:
- 0.1% n-dodecyl-beta-D-maltoside (DDM) — MS-compatible detergent
- 1 mM dithiothreitol (DTT) — reducing agent
- 100 mM ammonium bicarbonate (ABC) — buffering agent
- 10% DMSO
- Chemical decrosslinking compound (e.g., 1 mM 3,4-diaminobenzoic acid)
The decrosslinking agent is present during the critical extraction and lysis phase, not added as a separate post-extraction step.
Step 3 – Chemical-Aided Decrosslinking (Core Step)
The compound-containing buffer is incubated at the target temperature — typically 70–90°C — for 90–120 minutes. Key mechanistic points:
- Acts on crosslinked substrates catalytically: once substrate crosslinks are depleted, adding more reagent yields no further benefit
- Enables reduced-temperature protocols (70°C vs. 90°C), which is particularly valuable in nanodroplet workflows where evaporation is difficult to control
- Not all crosslink types respond equally — recent evidence points to cysteine thiol-based crosslinks as particularly amenable to chemical reversal
Step 4 – Alkylation, Digestion, and Purification
Post-decrosslinking steps include:
- Alkylate cysteines with chloroacetamide (CAA) to prevent disulfide reformation
- Digest with trypsin/Lys-C for bottom-up proteomics
- Purify via magnetic bead-based or silica column methods
For nucleic acid workflows, magnetic bead purification is standard. The Manta system from Cambrian Bioworks is designed for this stage: a compact, CE-IVD certified platform that processes 1–32 FFPE samples per run using pre-filled cartridges, with extraction completing in approximately 30 minutes after lysis. Its magnetic bead-based purification supports consistent DNA recovery for NGS, qPCR, and mutation analysis.
Step 5 – Quality Assessment and Downstream Application
Assess decrosslinking success by measuring:
For proteomics:
- Precursor and protein identification counts
- Coefficient of variation (CV) across replicates
- Recovery of specific peptide classes (e.g., cysteine-containing peptides)
For nucleic acids:
- Fragment size distribution and DIN score (>2.05 required for adequate NGS performance)
- Amplifiability (Ct values, delta Cq <5 for sequencing-grade DNA)
- Library complexity (true unique coverage)
Comparing these metrics against a thermal-only control validates whether chemical decrosslinking delivered improvement.
Evaluating Chemical Decrosslinking Agents: What the Research Shows
Researchers at Pacific Northwest National Laboratory systematically screened chemical compounds designed to catalyze decrosslinking in LCM-FFPE spatial proteomics using the nanoPOTS workflow on 100 µm × 100 µm regions of 10 µm thick human pancreatic FFPE tissue sections. Note: This is a 2026 preprint, not yet peer-reviewed.
Compound Panel and Selection Criteria
The team selected four compounds based on aqueous solubility, mass spectrometry compatibility, and structural analogy to known organocatalysts:
- 3,4-diaminobenzoic acid (3,4-DABA) — Compound A
- 2-amino-5-methylbenzoic acid — Compound B
- o-phenylenediamine — Compound C
- Bismuth tribromide — excluded due to insolubility
Key Outcomes at 90°C
| Compound | Precursor ID Change | Protein ID Change | CV (%) |
|---|---|---|---|
| Standard (thermal only) | Baseline | Baseline | 20% |
| 3,4-DABA | +14% | +6% | 13% |
| 2-Amino-5-methylbenzoic acid | +13% | +5% | 14% |
| o-Phenylenediamine | -17% | -8% | 17% |
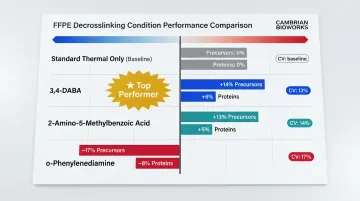
3,4-DABA yielded the best overall coverage with the lowest variability, making it the leading candidate for integration into spatial proteomics workflows.
Cysteine-Specific Decrosslinking Mechanism
Cysteine-containing precursors were significantly more abundant in compound-treated samples versus thermal controls. This points to 3,4-DABA selectively reversing thiol-specific cysteine crosslinks — a direct mechanism for improving recovery of the many functionally and clinically relevant proteins that carry cysteine residues.
Yet no significant differences were observed in formyl-lysine or methylene-bridge modifications on tryptophan/tyrosine. The improvement is selective: it targets specific crosslink classes rather than globally stripping formaldehyde modifications.
These mechanistic findings set the stage for the next question — what processing conditions extract the most from this selectivity?
Temperature and Time Optimization
At 70°C (reduced temperature):
- 3,4-DABA increased precursor identifications by 19% over the 70°C thermal-only control
- Protein identifications increased by 24% (from 2,639 to 3,283 proteins)
- This temperature reduction minimizes nanodroplet evaporation, a key practical advantage
Incubation time:
- Tested at 30, 60, 90, and 120 minutes
- Highest yield at 120 minutes (averaging 28,772 precursors and 3,985 proteins)
Concentration:
- Tested at 1 mM, 2.5 mM, 5 mM, and 7.5 mM
- No significant difference above 1 mM (plateau effect due to substrate-limited kinetics)
- 1 mM selected as optimal working concentration
Applications in Spatial Proteomics and Cancer Diagnostics
Spatial Proteomics: Near Single-Cell Resolution in FFPE
Laser capture microdissection (LCM) coupled with mass spectrometry enables protein mapping at near single-cell spatial resolution within intact tissue sections. FFPE compatibility is essential here because fresh-frozen tissue is rarely available for archival clinical samples.
The optimized 3,4-DABA protocol applied to a pancreatic ductal adenocarcinoma (PDAC) tissue section identified approximately 3,980 proteins across five distinct cell populations:
| Cell Population | Key Markers Identified |
|---|---|
| Acinar cells | PRSS1, PRSS2, CLPS, CPB1, CELA3A |
| Cancer/Adenoma cells | S100P, MYH14 |
| Dysplastic duct cells | CEACAM7 |
| Proximal stroma | Stromal markers |
| Distal stroma | Stromal markers |
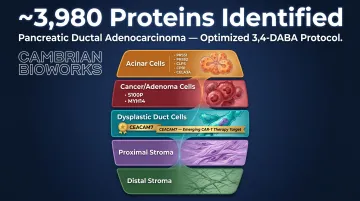
The identification of CEACAM7 in dysplastic ductal cells is particularly noteworthy. Independent research published in Clinical Cancer Research validated CEACAM7 as an effective CAR-T cell therapy target in PDAC, with preclinical models demonstrating tumor remission in patient-derived xenografts. That a therapeutic target of this caliber surfaced from archival tissue — using chemistry unavailable a decade ago — underscores what improved FFPE decrosslinking makes possible at spatial resolution.
Nucleic Acid Extraction and Clinical NGS Quality
The same crosslink chemistry that limits protein recovery also degrades nucleic acids. Organocatalytic removal of formaldehyde adducts from RNA and DNA bases — first described by Karmakar et al. in 2015 — directly informs nucleic acid-focused FFPE workflows.
FFPE-derived sequencing libraries show approximately 50% lower true unique coverage compared to fresh-frozen libraries, with several quality barriers:
- Average fragment sizes in the 300–400 bp range (vs. longer fragments in fresh-frozen)
- DIN scores often below the >2.05 threshold required for adequate NGS performance
- Amplifiability ratios as low as 5% in heavily degraded samples
Chemical decrosslinking improves recovery of amplifiable nucleic acid fragments long enough for library construction — often the difference between a failed and successful sequencing run for archival samples.
Clinical and Translational Implications
FFPE archives in hospitals span decades and are accompanied by pathology reports, diagnoses, and long-term clinical outcomes. Efficient chemical decrosslinking makes it feasible to extract meaningful proteomic and genomic data from these repositories at scale, enabling:
- Retrospective biomarker discovery in cohorts with known treatment outcomes
- Therapeutic target identification (e.g., CEACAM7 in PDAC) from archived tumor samples
- Mutation profiling for precision oncology when fresh tissue is unavailable
- Spatial molecular mapping of tumor microenvironment and immune infiltrates
For rare tumour types — where prospective fresh-frozen collection is logistically impossible — archived FFPE blocks may be the only available source of tumour material. Reliable decrosslinking converts those blocks from storage liabilities into actionable research assets.
How Cambrian Bioworks Can Help
Cambrian Bioworks provides automated, clinically validated nucleic acid extraction solutions designed specifically for FFPE sample workflows. The Manta automated extraction platform holds CE-IVD certification under Regulation (EU) 2017/746 and CDSCO approval, making it suitable for clinical diagnostic labs requiring reliable, regulatory-compliant DNA extraction at scale.
Key Advantages for FFPE Workflows
Purification that starts where decrosslinking ends:
Manta uses magnetic bead-based technology to purify DNA following lysis and decrosslinking, processing FFPE samples in approximately 28 minutes after lysis. This automated workflow reduces variability in the purification step, where inconsistency most impacts final nucleic acid quality for NGS, qPCR, and spatial omics applications.
Practical benefits for archival FFPE samples:
- No batching pressure: Process 1–32 samples per run as needed
- Minimal hands-on time: Pre-filled cartridge system reduces manual steps
- High-quality DNA output: Consistent purity ratios (A260/280 ~1.8–2.0), DIN scores suitable for sequencing, delta Cq values <5
- Compatible with diverse FFPE inputs: Validated across tumor types and specimen sites, including challenging over-fixed archival blocks

In use at major oncology centers:
Manta is deployed across 67+ labs globally. Tata Memorial Hospital and ACTREC (Advanced Centre for Treatment, Research and Education in Cancer) use the platform for oncology studies, mutation analysis, and NGS workflows from FFPE blocks.
For labs working with limited or archival FFPE material, pairing an optimised chemical decrosslinking protocol with automated extraction reduces the risk of sample loss at the purification stage — where manual handling introduces the most variability.
Frequently Asked Questions
What is the difference between chemical decrosslinking and heat-induced antigen retrieval in FFPE samples?
Heat-induced antigen retrieval (HIAR) uses high temperature (typically 95–100°C in citrate or EDTA buffer) to expose epitopes for antibody binding in immunohistochemistry. Chemical decrosslinking adds catalytic compounds to enzymatically reverse formaldehyde crosslinks during protein or nucleic acid extraction for mass spectrometry or sequencing. Chemical approaches offer more complete crosslink reversal and are designed for molecular analysis rather than antibody-based detection.
Why does formalin fixation create crosslinks, and why are they difficult to reverse?
Formalin reacts with nucleophilic groups on proteins (lysine, cysteine, histidine) and nucleic acids to form methylol adducts and methylene bridges, creating intra- and inter-molecular covalent bonds. These crosslinks are thermally stable — high-temperature treatment reverses only heat-labile methylol adducts, leaving the more stable methylene bridges intact and requiring catalytic chemical agents for complete reversal.
What makes 3,4-diaminobenzoic acid effective as a decrosslinking agent?
3,4-diaminobenzoic acid is particularly effective at reversing thiol-specific cysteine crosslinks, delivering significantly higher recovery of cysteine-containing peptides. It works catalytically, is aqueous-soluble, and integrates cleanly into mass spectrometry workflows — achieving the lowest coefficient of variation (13%) among agents tested in recent spatial proteomics research.
Does chemical decrosslinking also improve RNA and DNA extraction from FFPE tissues?
Yes. Organocatalytic removal of formaldehyde adducts from nucleic acid bases is the conceptual foundation for chemical decrosslinking, and the evidence is clear: chemistry-based extraction yields 7–25-fold more amplifiable nucleic acids than heat-only methods, directly benefiting NGS, qPCR, and microarray workflows on archival FFPE samples.
What downstream applications benefit most from improved FFPE decrosslinking?
The highest-impact applications are:
- Spatial proteomics: LCM-MS for near single-cell protein mapping
- Bottom-up proteomics: Biomarker discovery from limited tissue
- NGS: Whole-exome sequencing, targeted oncology panels, and RNA-seq on archival samples
- Clinical diagnostics: Workflows where DNA/RNA quality directly determines variant calling accuracy
How do I choose between thermal-only and chemical-aided decrosslinking for my FFPE workflow?
Thermal-only methods are adequate for bulk, high-input samples with standard downstream applications. Choose chemical-aided decrosslinking when working with:
- Low-input or single-cell spatial proteomics
- Heavily fixed archival tissue
- RNA applications sensitive to thermal degradation
If you're consistently seeing a 10–15% proteome gap or poor DNA amplifiability, chemical decrosslinking is the right next step.


