The core challenge is straightforward: traditional scRNA-seq requires fresh or snap-frozen tissue, demanding immediate sample processing and precise logistical coordination. This limits researchers to prospective study designs, excluding rare diseases, retrospective cohorts, and samples with known clinical outcomes already documented in patient records.
The breakthrough came through probe-based chemistries and specialized nuclei isolation protocols. These technologies are transforming FFPE tissue from a liability into an asset, unlocking decades of archived clinical material for high-resolution single-cell discovery without the constraints of real-time sample collection.
TLDR
- FFPE tissue yields fragmented RNA — incompatible with conventional poly(dT)-based scRNA-seq
- Probe-based methods like 10x Flex chemistry bypass fragmentation by targeting short RNA sequences
- Nuclei isolation produces higher-quality single-nucleus suspensions than whole-cell dissociation
- Data quality depends more on fixation and dissociation protocols than on block age
The Hidden Goldmine: Why FFPE Tissue Archives Matter for Single-Cell Research
Pathology departments routinely preserve tissue from surgical and biopsy procedures as FFPE blocks, creating repositories that span years to decades. Major cancer centres maintain FFPE archives containing millions of specimens, representing the single largest source of clinically annotated human tissue worldwide.
Strategic advantages of FFPE archives:
- Select samples retroactively by defined clinical criteria — tumor subtype, mutation status, treatment history — enabling statistically powered cohort design
- Access rare disease populations that no single prospective study could realistically recruit
- Correlate molecular profiles directly with long-term outcomes using samples that already have complete follow-up data
FFPE workflows also democratise single-cell access. Fresh-tissue studies require real-time coordination between clinicians, researchers, and labs — a logistical barrier that has historically confined single-cell work to well-resourced academic centres. FFPE-compatible methods work with existing archived material, making single-cell discovery viable for laboratories worldwide.
The Core Challenge: How Formalin Fixation Damages RNA and Why Standard scRNA-seq Fails
Mechanism of RNA Degradation
Formalin fixation creates protein-RNA and protein-protein crosslinks, fragmenting RNA into short pieces. High heat during paraffin embedding further degrades integrity. The result: FFPE RNA typically shows poor DV200 scores—the percentage of RNA fragments above 200 nucleotides—a key quality metric for sequencing suitability.
Why Conventional Protocols Fail
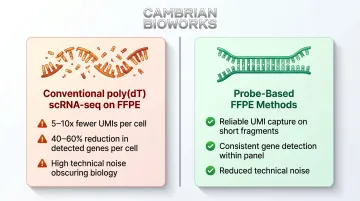
Methods like 10x 3' and SMART-seq2 rely on poly(dT) probes capturing intact poly-A tails on full-length mRNA for cDNA synthesis. Fragmented FFPE RNA lacks intact poly-A tails at usable density, causing dramatic underperformance:
- 5-10x fewer unique molecular identifiers (UMIs) per cell
- 40-60% reduction in detected genes per cell
- High technical noise obscuring biological variation

Whole-Cell Dissociation Adds Further Failure
Unlike frozen tissue, deparaffinised FFPE sections don't dissociate cleanly into single-cell suspensions. Enzymatic dissociation produces:
- Partially intact cells with leaking cytoplasm
- Free nuclei from lysed cells
- Tissue debris interfering with microfluidic capture
This mixture causes inaccurate cell counting, doublet inflation, and poor droplet loading in microfluidic devices.
Nuclear RNA as the Solution
Isolating nuclei sidesteps both problems. When cells dissociate incompletely from FFPE, cytoplasmic RNA leaks and degrades further — but nuclear RNA remains protected within an intact membrane, providing consistent input. Studies show nuclei extraction removes tissue debris, reducing ambient RNA contamination in droplet barcoding workflows.
FFPE nuclei isolation also captures cell populations that fresh dissociation misses. In fresh protocols, epithelial cells undergo apoptosis-induced stress artifacts during processing, and collagen-embedded fibroblasts are chronically underrepresented.
FFPE fixation locks spatial integrity in place before nuclei are isolated, preserving these populations for more accurate downstream profiling.
Modern FFPE Single-Cell Workflows: Probe-Based Solutions and Nuclei Isolation Strategies
The Probe-Hybridisation Paradigm Shift
Probe-based detection targets short (~50 bp) RNA sections rather than requiring full-length integrity. 10x Flex (Fixed RNA Profiling) chemistry exemplifies this approach—probes hybridise to fragmented RNA, and probe molecules (not RNA itself) are barcoded and sequenced.
This enables reliable detection on degraded samples, though coverage is limited to the probe panel. The Human Transcriptome v1.1.0 panel targets 18,526 genes with 53,957 probes, excluding 397 genes by default — including highly polymorphic genes like MHC class II.
snPATHO-seq: The Dedicated FFPE Workflow
snPATHO-seq represents the most mature dedicated FFPE single-nucleus method, with key steps:
- Deparaffinisation: Xylene washes remove paraffin
- Rehydration: Serial ethanol gradient
- Enzymatic dissociation: Liberase/Collagenase D digestion
- Nuclear lysis: Ez lysis buffer treatment
- Sequential filtration: 70 µm then 40 µm mesh
- Library preparation: 10x Flex chemistry
This dedicated nuclei isolation delivers more consistent results than simpler workflows like 10x scFFPE, which produced mixed cell/nuclei/debris suspensions in dense tissues like breast cancer. That distinction matters when working with architecturally complex samples where clean nuclear capture directly affects data quality.
snRandom-seq and Total RNA Strategies
snRandom-seq uses random primers rather than poly(dT) to amplify all RNA fragments, then polyadenylates the resulting cDNA. This total RNA approach offers:
- Broader transcriptomic coverage including non-coding RNA
- Higher UMI and gene counts per nucleus versus probe-based methods
- Requires specialised chemistry, now commercialised and automated
Studies show snRandom-seq achieved median detection of 1,635 genes in mouse brain FFPE samples, demonstrating viability across tissue types.
Why Single-Nucleus Over Single-Cell for FFPE
Single-nucleus approaches are generally preferred because:
- Nuclei are mechanically stable for isolation from fixed tissue
- Uniform input for droplet capture reduces technical variability
- Better representation of tissue-resident cell types in dense tissues (brain, liver, kidney, tumours)
- High concordance with scRNA-seq for most cell type markers
The case for nuclei isolation extends beyond practicality — it also opens the door to multi-modal experimental designs.
Multi-Modal Integration Advantage
FFPE samples prepared for snRNA-seq can be paired with FFPE spatial transcriptomics (such as 10x Visium FFPE) on sections from the same block. This enables:
- Single-nucleus gene expression profiled alongside spatially resolved transcriptomics
- Spatial deconvolution using single-nucleus data to assign cell types to Visium spots
- Richer tissue characterisation than either modality achieves independently
Getting It Right: Practical Sample Preparation and Quality Control for FFPE scRNA-seq
Fixation Quality Matters Most
Neither block age alone nor DV200 score reliably predicts success. The adequacy of original formalin fixation is critical:
- Under-fixation: RNA degrades rapidly post-resection
- Over-fixation: Several days in formalin (versus standard ~24 hours) over-crosslinks nucleic acids and protein, impairing lysis and probe hybridisation
- Pathologist review: H&E morphology assessment identifies necrotic or poorly preserved areas to exclude before sequencing
Tissue Input and Cellularity
Dense tissues (high-cellularity lung carcinoma) require less input than low-cellularity tissues. Published studies validate:
- Single tissue microarray (TMA) cores of 1-1.5 mm diameter work for high-cellularity samples
- Typical section thickness: 25-30 µm for snPATHO-seq
- Optimise section number based on expected cell yield
Enzymatic Dissociation Optimisation
Tissue type drives protocol variation:
- Fibrous/collagen-rich tissues (breast) require longer incubation or higher-speed homogenisation
- Automated liquid handlers like Cambrian Bioworks' Beluga standardise multi-step enzymatic digestion, washing, and resuspension
- Automation reduces manual variability in nuclei isolation, where inconsistent technique most directly affects downstream yield
Once dissociation is complete, QC checkpoints before library preparation determine whether the sample is ready for microfluidic loading.
Quality Control Checkpoints
Assess each of the following:
- Nuclei yield and viability: Use cell counter with AO/PI staining to distinguish intact from damaged nuclei
- Debris contamination: Visual check under phase-contrast microscopy
- FACS sorting: DAPI staining enriches intact nuclei, removes debris, and provides accurate counts for microfluidic loading; DAPI is fully compatible with FFPE nuclei and does not impair sequencing quality

What Researchers Are Discovering: Applications of FFPE scRNA-seq in Cancer and Beyond
Reshaping Tumour Microenvironment Research
FFPE scRNA-seq enables retrospective cohort studies correlating single-cell profiles with known clinical outcomes, mutation status, and treatment response. Researchers identified novel tumour cell subpopulations, such as multi-ciliated neoplastic cells in breast cancer IDC, that would have been missed in small prospective fresh-tissue studies.
Validation Through Concordance
Benchmarking studies show major cell type proportions, canonical marker expression, and copy number variation profiles from FFPE match fresh/frozen tissue. This gives researchers confidence that insights from archival samples are biologically valid, though lowly expressed genes may show slightly less correlation than highly expressed canonical markers.
That biological reliability is precisely what makes FFPE workflows valuable beyond cancer research, where archival collections span decades of well-annotated clinical histories.
Beyond Oncology
FFPE single-nucleus workflows are expanding into:
- Neurological disease studies, including archived glioma samples profiled via snRandom-seq
- Inflammatory and autoimmune disease research, where prospective fresh-tissue collection is often logistically constrained
- Rare disease biobanks, where ethical or practical barriers make prospective sampling impossible
For labs already sitting on years of well-annotated archival tissue, this means existing collections can now yield single-cell resolution data — without a single new biopsy.
Frequently Asked Questions
Can you do single-cell RNA sequencing on FFPE tissue?
Yes, FFPE-compatible scRNA-seq is well-established using probe-based methods like 10x Flex chemistry or total RNA approaches like snRandom-seq. Nuclei isolation rather than whole-cell dissociation is the preferred sample preparation strategy for most tissue types.
Why is RNA quality poor in FFPE samples compared to fresh or frozen tissue?
Formalin fixation creates protein-RNA crosslinks and fragments RNA molecules, while high heat during paraffin embedding further degrades integrity. FFPE RNA is predominantly short fragments that conventional poly(dT)-dependent methods cannot capture.
What is the difference between scRNA-seq and snRNA-seq, and which is better for FFPE?
scRNA-seq sequences RNA from whole cells while snRNA-seq sequences nuclear RNA. For FFPE, single-nucleus approaches are preferred because intact nuclei are more consistently isolatable from fixed tissue, and nuclear RNA reliably reflects cellular transcriptional states.
How does the age of an FFPE block affect single-cell RNA sequencing results?
Block age alone is not the primary determinant of data quality—original fixation quality matters more. Well-fixed blocks can yield usable data after several years of storage, though RNA quality does decline over time, especially beyond 5-8 years.
Which FFPE single-nucleus workflow gives the best gene coverage?
snRandom-seq offers broader transcriptomic coverage (total RNA including non-coding transcripts) while probe-based methods like snPATHO-seq/10x Flex offer reliable cell-type detection within a defined gene panel. The optimal choice depends on the research question and sample characteristics.
What tissue types work best for FFPE single-cell sequencing?
High-cellularity solid tumours (lung, breast, colon cancers) and tissues with clear cell-type heterogeneity have been most extensively validated. Fibrous or collagen-rich tissues require protocol optimisation for dissociation, and necrotic tissue regions should be excluded.


